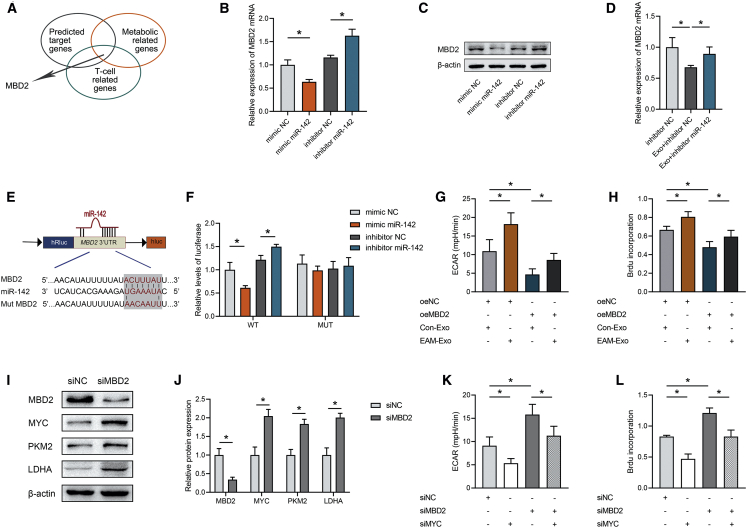
Figure 5.
Exosomal miR-142 Regulates CD4+ T Cell Glycolysis and Metabolic Reprogramming by Targeting MBD2
(A) Screening scheme for target genes that might contribute to the effects of T cell metabolic regulation mediated by miR-142. (B) Relative mRNA expression of the target gene MBD2 in CD4+ T cells treated with miR-142 mimic negative control (NC), miR-142 mimic, inhibitor NC, or miR-142 inhibitor. (C) Representative blots showing protein expression of the target MBD2 in CD4+ T cells treated with miR-142 mimic NC, miR-142 mimic, inhibitor NC, or miR-142 inhibitor. (D) Relative mRNA expression of MBD2 in CD4+ T cells after exosome incubation. (E) The matched base pairs in the binding region of miR-142 and the reporter plasmids containing the wild- or mutant-type MBD2 are shown. (F) Relative luciferase reporter activity of vectors carrying the luciferase gene and a fragment of MBD2 3′UTR with wild- or mutant-type miR-142 binding sites after cotransfection with miR-142 mimic NC, miR-142 mimic, inhibitor NC, or miR-142 inhibitor. (G) ECARs were measured in CD4+ T cells transfected with MBD2 and incubated with exosomes. (H) BrdU incorporation was calculated in CD4+ T cells transfected with MBD2 and incubated with exosomes. (I and J) Representative blots (I) and quantified data (J) showing protein expression of MBD2, MYC, and metabolic enzymes in CD4+ T cells transfected with siMBD2. (K) ECARs were measured in CD4+ T cells transfected with siMBD2 and siMYC. (L) BrdU incorporation was calculated in CD4+ T cells transfected with siMBD2 and siMYC. Data are presented as the means ± SEM (n = 6); ∗p < 0.05.