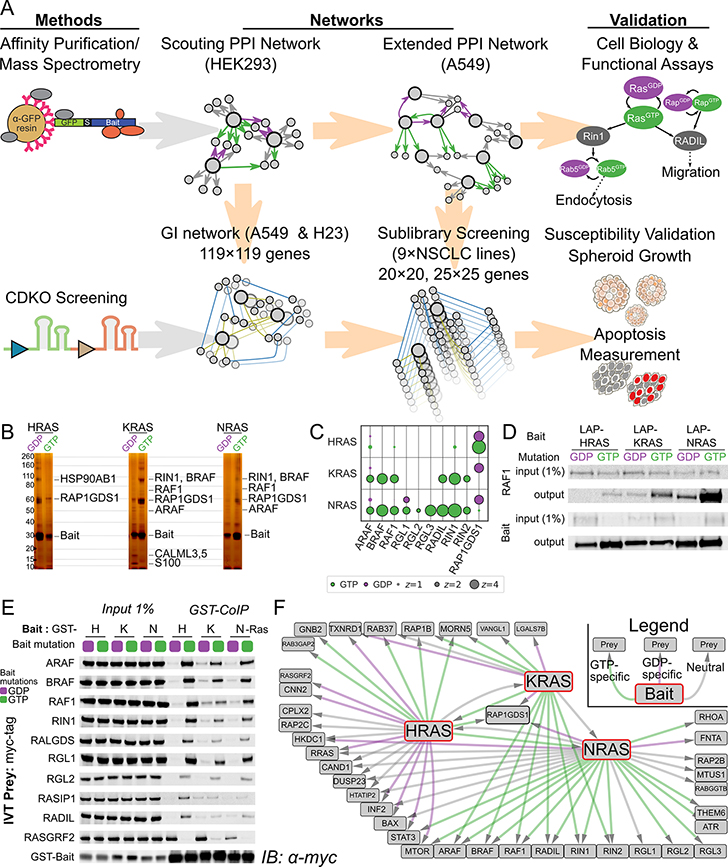
Figure 1. Multiomic Interaction Mapping Reveals Strong Selectivity of RAS Isoforms.
A Schematic outlining experimental approach. Top row: Proteins of known interest, expressed as GFP-tagged constructs, were used as bait for AP/MS to discover protein-protein interactions (PPIs). Proteins from the PPI network were selected for CRISPR Dual Knockout (CDKO) genetic interaction (GI) screening. sgRNA cassettes targeting these genes were transduced into Cas9-expressing KRAS-driven LUAD lines and assayed for enrichment or depletion after 21 days growth in culture. Relative depletion of single- and double-targeting cassettes was used to establish synergy or buffering between gene knockout phenotypes. GIs that could form the basis of new therapies were validated by both CDKO and AP/MS in other LUAD lines, as well as functional and growth assays. B Silver stain PAGE of tandem affinity purification of H-, K-, and NRAS from A549 cells shows distinct GTP-specific PPIs. Known effectors are marked. C Bubble plot of 6 AP/MS experiments with GTP- and GDP-locked mutant GTPases as baits (rows), showing the enrichment of selected preys (columns). Dark borders indicate FDR < 0.05. D Western blot of co-immunoprecipitation reactions from A549 cells expressing tagged H, K, or NRAS with mutations locking them in their GTP or GDP-bound forms, probing for endogenous RAF1. HRAS binds significantly less RAF1 than does either other RAS. E Cell-free co-immunoprecipitation of selected prey proteins by recombinant purified mutant H, K, and NRAS, imaged by immunoblot. Negligible RAS isoform specificity is observed. F Network diagram showing PPIs of RAS proteins and selected prey proteins. Green and purple lines indicate PPIs from GTP- and GDP-locked preys respectively; gray lines indicate that the PPI is not nucleotide-state specific. All diagrammed interactions have FDR < 0.05 (see Methods)