Fig. 6.
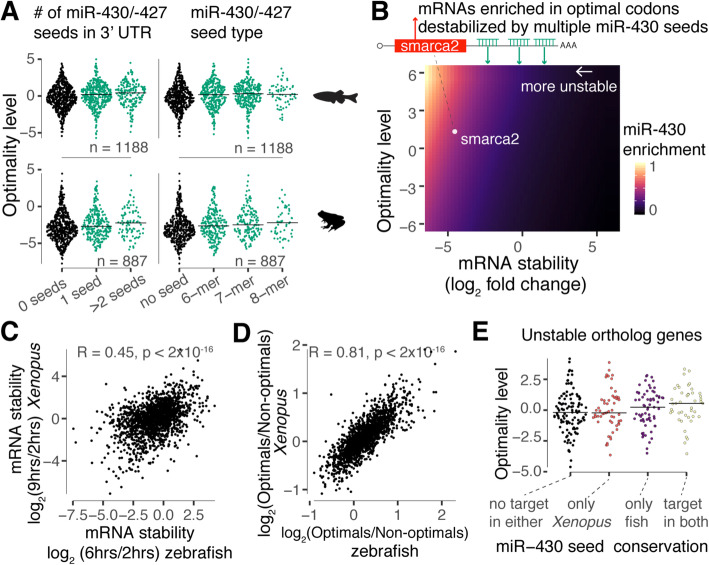
MicroRNAs antagonize codon optimality effect on mRNA stability during the MZT. a Sinaplot showing the codon optimality distribution (Table S5) for the top 1000 most unstable maternal genes during zebrafish and Xenopus MZT. The stability was defined based on the log2-fold change of early (2 h) vs late time points (fish = 6 h, Xenopus = 9 h) in fish (Additional file 1: Fig. S1a-b) and Xenopus [33]. The genes were divided into groups based on the numbers or seed type of miR-430/427. The content of optimal codons increases with the miR-430 regulation strength (p = 2 × 10−4 zebrafish, p = 2 × 10−3 Xenopus, linear regression). The p value was obtained by comparing the difference in the mean level of optimal codons (PLS1) between genes with and without miR-430/427 sites using a linear model. b Heatmap of miR-430 enrichment in the 3′UTR as a function of mRNA stability and codon optimality level. The miR-430 enrichment was estimated with a generalized linear model. Unstable mRNAs enriched in optimal codons temp to contain miR-430 sites (e.g., smarca2). c Scatter plot comparing the RNA stability, during MZT, of ortholog genes in zebrafish (Additional file 1: Fig. S1a-b) and Xenopus [33]. d Scatter plot comparing the content of optimal codons in zebrafish and Xenopus for ortholog genes [22]. e Sinaplot showing the codon optimality distribution for unstable mRNAs in panel a that are orthologs (n = 280). Messenger RNAs were divided into four categories according to the presence or absence of miR-430/-427 seeds in both species. The orthologous mRNA with miR-430/-427 seeds in both species are the most enriched in optimal codons (p = 0.0868, one-way ANOVA test)