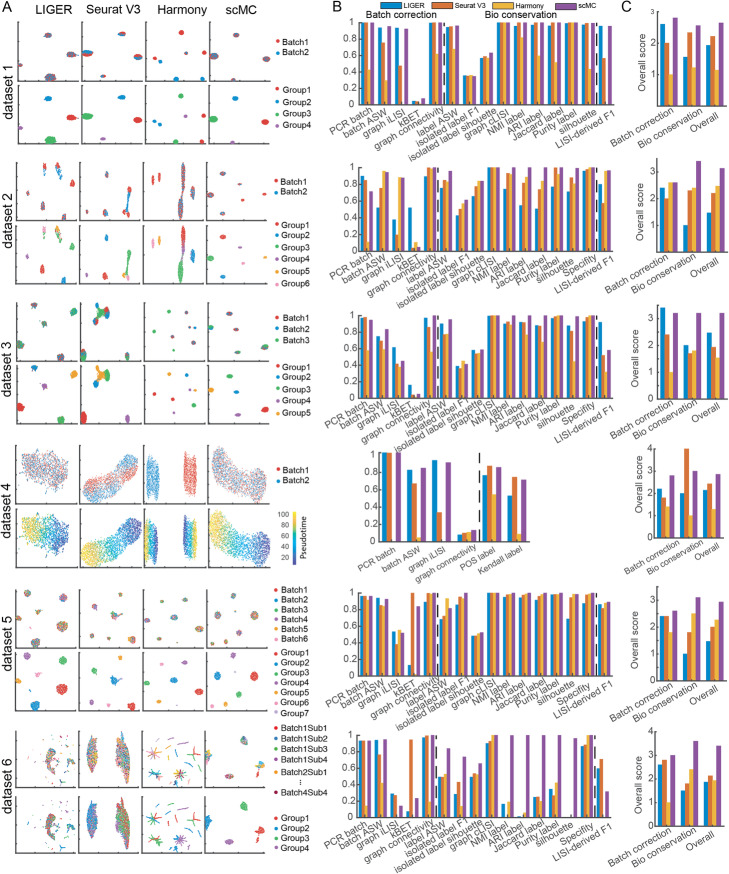
Fig. 2.
Comparisons of scMC against other methods on six simulation scenarios. a UMAP visualization of the corrected data from LIGER, Seurat V3, Harmony, and scMC. For each dataset, cells are colored by batch labels (top row) and gold standard cell labels or pseudotime (bottom row). b Evaluation of integration methods using 16 metrics in two categories for measuring batch effect removal (i.e., Batch correction) and biological variation conservation (i.e., Bio conservation). LISI-derived F1 score is a summarized metric assessing both batch mixing and cell type separation. The consistency between the inferred pseudotime from the corrected data and the gold standard pseudotime is computed using two metrics POS and Kendall rank correlation coefficient. c Comparison of the overall scores among different methods calculated based on batch effect removal metrics, biological variation conservation metrics, and both batch effect removal and biological variation conservation metrics