Fig. 5.
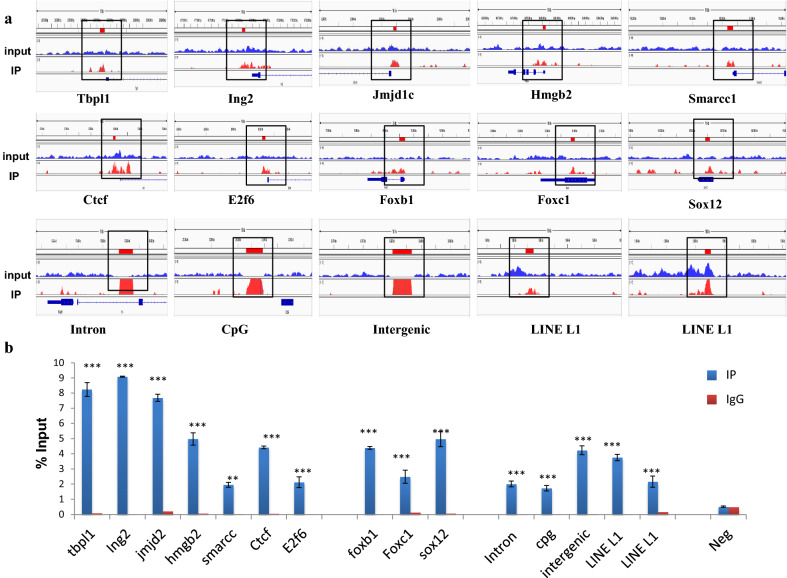
Validation of H1T2 ChIP-sequencing data by ChIP PCR analysis of representative genes bound to the linker histone H1T2. a Screenshots of H1T2 ChIP peaks displayed from Integrative Genomics Viewer. Shown are 15 different genomic regions associated to H1T2. The y-axis unit is reads per million (rpm).The x-axis represent the genomic positions in base pairs as follows: Chr1:23977211–23977579; Chr16:47717217–47717457; Chr20:22882800–22883032; Chr16:36080014–36080219; Chr8:118206347–118206725; Chr19:37599804–37600003; Chr6:42092121–42092366; Chr8:75994988–75995401; chr17:33949714–33950006; Chr3:147864851–147865149; Chr5:173429327–173430483; Chr1:213008609–213009881; Chr15:33478042–33479362; Chr5:1682868–1683456; ChrY:1588659–1589079. b Bar graphs showing H1T2 ChIP-PCR results represented as % of input. The first seven regions represents the H1T2 occupancy in transcription factors and the next group of three regions represent the H1T2 bound developmental genes while the remaining regions shows the preferential binding of H1T2 to the repeat elements. A negative control region (specific regions of chr 15) was also included for ChIP PCR, where no peaks were present according to the data analysis and visualization. Results are from three independent experiments and error bars represent standard deviation. *shows comparison with IgG control and # shows comparison with IP + pep group. ***(P ≤ 0.001), **(P ≤ 0.01), *(P ≤ 0.05)