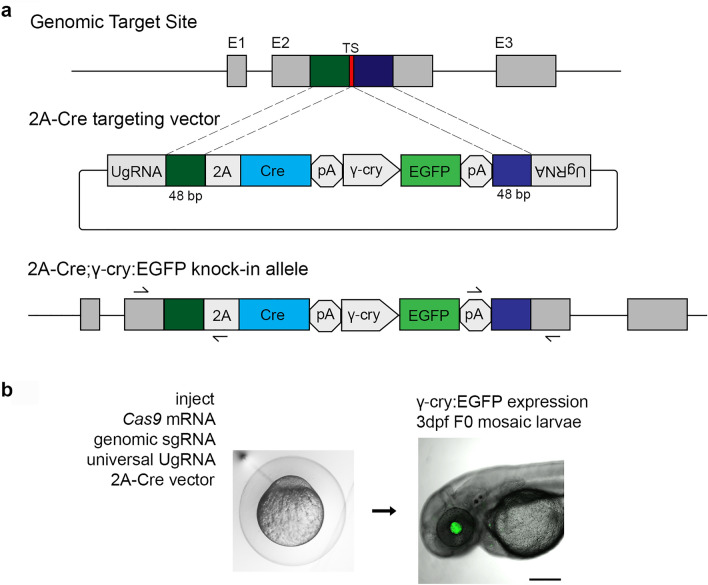
Figure 1.
CRISPR/Cas9 short homology directed targeted integration strategy for efficient recovery of Cre knock-in alleles. (a) Schematic of the genomic target site, Cre donor vector, and final knock in allele. A CRISPR sgRNA (red) was chosen in the coding sequence. 48 bp upstream (dark green) and 48 bp downstream (dark blue) of the cut site are included in the donor vector as homology arms. The donor vector has UgRNA sequences flanking the homology arms for in vivo liberation of the knock in cassette. The knock-in cassette contains an in frame 2A-Cre for expression of Cre recombinase, and a secondary marker driving γ-cry:EGFP expression in the lens. Arrows indicate PCR primers for 5′ and 3′ genome/vector junction analysis. (b) 1 cell stage zebrafish embryos are injected with Cas9 mRNA (300 pg), genomic sgRNA (25 pg), UgRNA (25 pg), and 2A-Cre targeting vector (10 pg). At 3 days post fertilization, embryos expressing EGFP in the lens are selected and raised to adulthood. TS genomic sgRNA target site, UgRNA universal short guide RNA target site, γ-cry gamma crystallin promoter, EGFP green fluorescent protein, pA transcription termination and polyadenylation sequence.