Fig. 6. The 2c-like transcriptome in the absence of SMCHD1 is partially dependent on Dux.
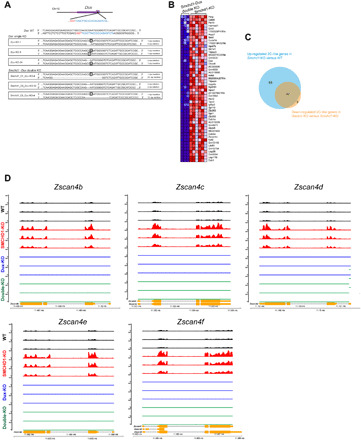
(A) The small guide RNA (gRNA) targeting region is immediately downstream of the start codon (ATG) of the Dux gene. Sanger sequencing confirmed frameshift mutations. Sequences targeted by the gRNA are in blue, and the PAM (protospacer adjacent motif) sequence is shown in red. Biallelic frameshift mutation was shown for each clone. The gRNA was applied in WT and in SMCHD1 KO ES cells to obtain the Dux single-knockout and Smchd1/Dux double-knockout ES cell clones, respectively. (B) A heatmap indicates the differentially expressed 2c-like genes between Dux/Smchd1 double-knockout (n = 3 clones) and Smchd1 single-knockout (n = 3 clones, two replicates each) ES cells. The blue color indicates genes no longer up-regulated in the double knockouts. (C) The pie chart shows that 47 of the 136 single-KO up-regulated 2c-like genes were no longer up-regulated in the absence of Dux in the Smchd1/Dux double KO. (D) RNA-seq tracks generated by the GVIZ package across the Zscan4 gene family members in WT ES cells (black), Smchd1 KO ES cells (red), Dux KO ES cells (blue), and Smchd1 plus Dux (double-KO) ES cells (green).
