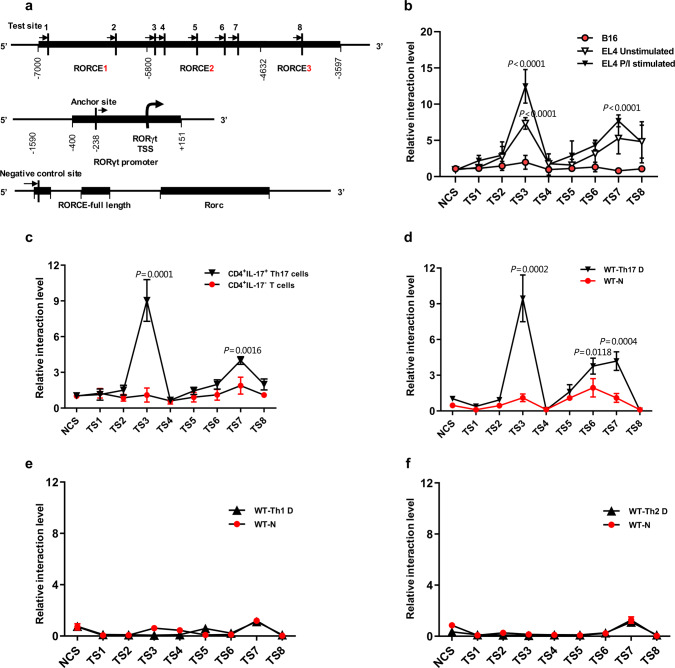
Fig. 5. 3C analysis of the RORCE region.
a Map of the investigated interactions. Since we found NlaIII to be the only restriction enzyme that allowed sufficiently high-resolution analysis of RORCE, NlaIII was finally used to digest the RORCE region. The positions of NlaIII sites (vertical lines) in RORCE used in this 3C analysis are indicated as test sites 1 to 8 (TS1 to TS8). The NlaIII site in RORγt gene promoter was taken as the anchor site (AS), from which possible interactions with other regions of RORCE were quantified. The primers used for 3C-qPCR are indicated by simple black arrows. The negative control site (NCS) is another NlaIII site upstream of RORCE, which does not interact with the AS. b–f Quantitative analysis of a 3C-qPCR assay assessing the RORCE region. The data represent relative interaction frequencies between AS and the other tested sites (TS1 to TS8) in the RORCE region. Relative interaction frequencies were determined by qPCR relative to standard curves as described in the “Methods” section. Data points represent the mean ± SEM for PI-stimulated EL4 (b, n = 4 independent experiments), FACS-purified CD4+IL17+ Th17 (c, n = 5 independent experiments), Th17-polarized (d, n = 5 independent experiments), Th1-polarized (e, n = 5 independent experiments), and Th2-polarized cells (f, n = 5 independent experiments) from WT mice. Th17 D, Th1 D, and Th2 D represent Th17, Th1, or Th2 differentiation under the appropriate polarization conditions, respectively, and N represents naïve cells. All P-values are based on unpaired two-tailed Student’s t-test (b–d). Experimental mice were between 8 and 12 weeks of age, with no preference to gender and were maintained on a C57BL/6 background. Source data are provided as a Source Data file.