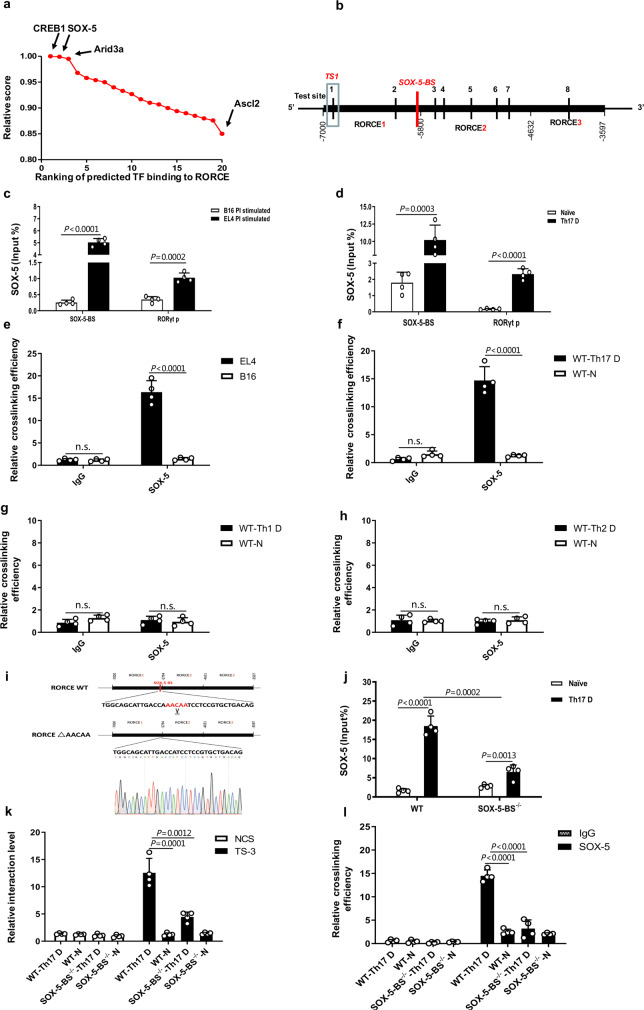
Fig. 6. ChIP-loop analysis of the RORCE region.
a The ranking of predicted transcriptional factors (TFs) capable of binding to RORCE was performed based on relative scores predicted with the JASPAR database (http://jaspar.genereg.net). b Red vertical line shows SOX-5 binding site (SOX-5-BS) in RORCE, which is very close to TS3. c, d A ChIP-qPCR assay was performed to determine the enrichment of SOX-5 in SOX-5-BS and RORγt promoter (RORγt p) in PI-stimulated EL4 cells (c) and WT Th17-polarized cells (d). e–h A ChIP-loop assay with an anti-SOX-5 antibody or control mouse IgG was performed to quantify relative interaction efficiency between the AS and TS3 mediated by SOX-5 in PI-stimulated EL4 (e), Th17-polarized (f), Th1-polarized (g), and Th2-polarized cells (h) from WT mice. i The sequence of SOX-5-BS (AACAA) in RORCE was deleted by using CRISPR/Cas9 technology (top). The knockout allele was confirmed by sequencing (bottom). j A ChIP-qPCR assay was performed to test the enrichment of SOX-5 in SOX-5-BS in Th17-polarized cells from SOX-5-BS-deficient (SOX-5-BS−/−) or WT mice. k, l Quantitative analysis of 3C-qPCR (k) and ChIP-loop assays (l) evaluating the interaction between TS3 and AS in Th17-polarized cells from SOX-5-BS-deficient or WT mice was performed. Th17 D, Th1 D, and Th2 D represent Th17, Th1, or Th2 differentiation under the corresponding polarization conditions, respectively, and N represents naïve cells. Mean ± SEM are shown, n = 4 independent experiments, unpaired two-tailed Student’s t-test (b–h, j–l). Experimental mice were between 8 and 12 weeks of age, with no preference to gender and were maintained on a C57BL/6 background. Source data are provided as a Source Data file.