Figure 2. Protein model illustrating impact of cyp51A mutations on binding of the frontline anti-Aspergillus triazole, voriconazole.
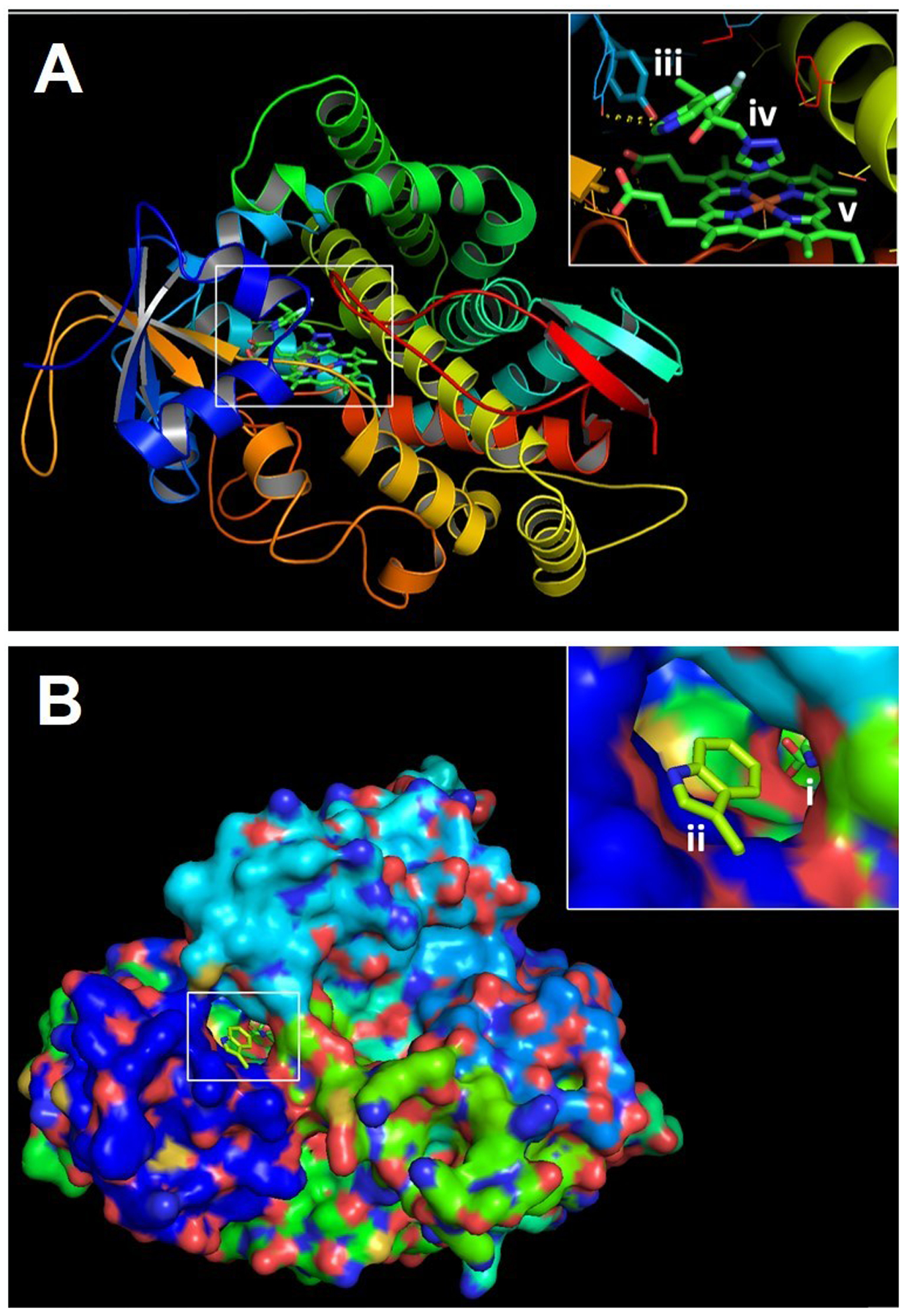
PyMOL 2 was used to create three dimensional models of the A. fumigatus Cyp51A protein using the previously resolved crystal structure of the A. fumigatus Cyp51B as a scaffold. A) Three-dimensional ribbon model of Cyp51A. Inset depicts the Y121 residue (iii) which stabilizes the interaction between voriconazole (iv) and heme (v). B) Three-dimensional model of the surface of the Cyp51A protein. Inset depicts the ligand access channel and entrance to the Cyp51A catalytic site which is occupied by both heme and voriconazole (i). Also depicted is the G54W amino acid substitution (ii) predicted to interfere with the binding of the long lipophilic sidechains of itraconazole and posaconazole.
