Figure 1.
Quantitative direct detection of viral SARS-CoV-2 RNA with Cas13a
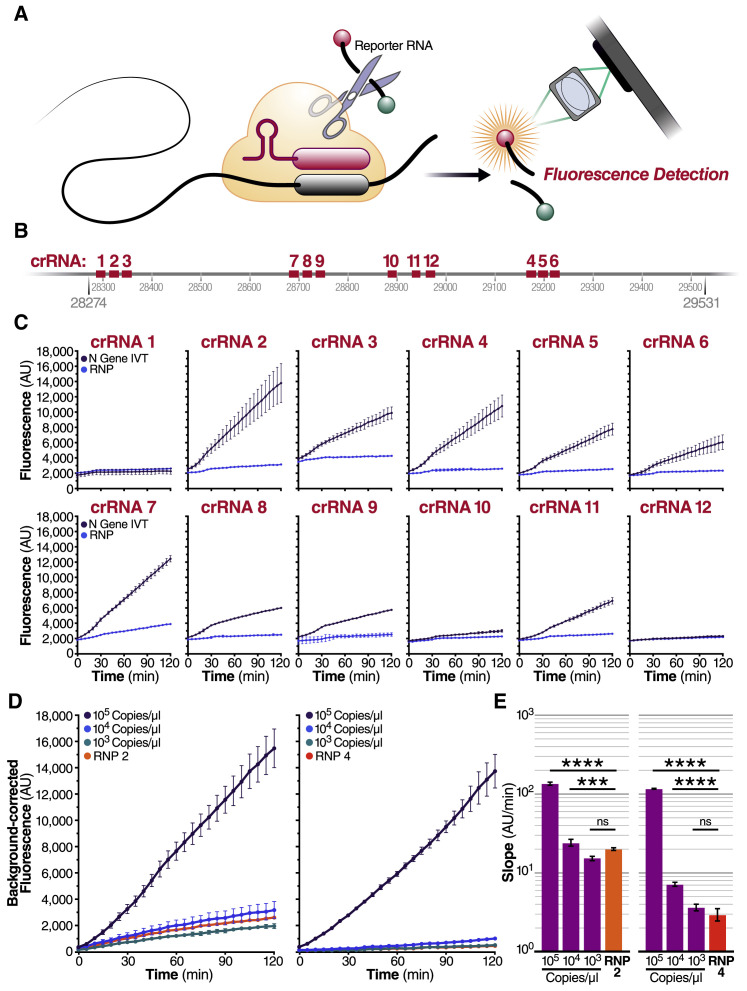
(A) Schematic of a Cas13a (beige)-crRNA (red) RNP complex binding target RNA (black), resulting in activation of the HEPN nuclease (denoted by scissors) domain. Upon target recognition and RNP activation, Cas13a indiscriminately cleaves a quenched-fluorophore RNA reporter, allowing for fluorescence detection as a proxy for Cas13a activation and target RNA.
(B) Schematic of the SARS-CoV-2 N gene, and the corresponding location of each crRNA spacer region.
(C) Cas13a RNPs made individually with each crRNA were tested against 2.89 × 105 copies/μL (480 fM) of SARS-CoV-2 IVT N gene RNA in a total 20 μL reaction volume. Background fluorescence by the individual RNP in the absence of target RNA is shown as “RNP.” Raw fluorescence values over 2 h is shown. Data are represented as mean ± standard deviation (SD) of three technical replicates. See also Figure S1A.
(D) Limit of detection of crRNA 2 and crRNA 4 was determined by testing 100 nM of each RNP individually against 105, 104, and 103 copies/μL of N gene IVT RNA. “RNP 2” and “RNP 4” represent no target RNA RNP alone controls. Background correction of fluorescence was performed by subtraction of reporter alone fluorescence values. Data are represented as mean ± standard error of the difference between means of three technical replicates. See also Figures S1B–S1D.
(E) Slope of the curve over 2 h from Figure 1D was calculated by performing simple linear regression to data merged from replicates and is shown as slope ± 95% confidence interval. Slopes were compared to the RNP alone control through an analysis of covariance (ANCOVA): ∗∗∗∗p < 0.0001, ∗∗∗p < 0.001, ns = not significantly higher than RNP control.