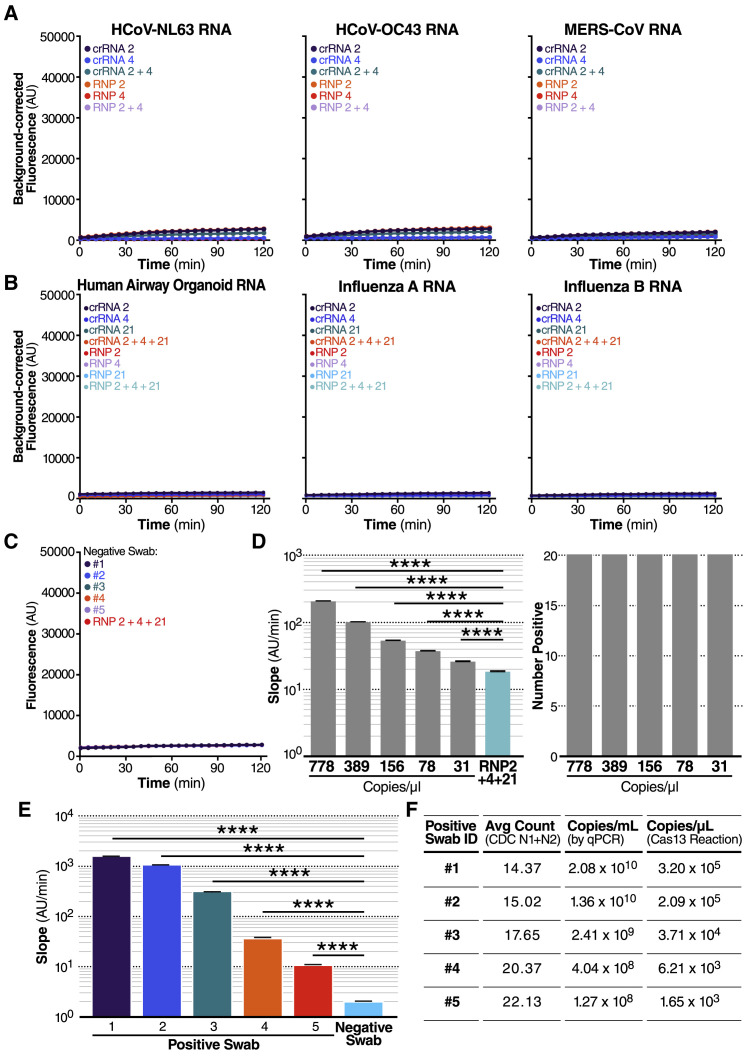
Figure 3.
Cas13a directly detects SARS-CoV-2 RNA in patient samples
(A) crRNA 2 and crRNA 4 were tested individually (100 nM total RNP concentration) and in combination (100 nM total RNP concentration: 50 nM each of RNP 2 and RNP 4) against RNA isolated from HCoV-NL63 viral supernatant (left) and HCoV-OC43 viral supernatant (center) or the IVT N gene RNA from MERS-CoV (right). No target RNA RNP alone controls are denoted as “RNP 2,” “RNP 4,” and “RNP 2+4.” Background correction of fluorescence was performed by subtraction of reporter alone fluorescence values. Data are represented as mean ± standard error of the difference between means of three technical replicates.
(B) crRNA 2, crRNA 4, and crRNA 21 were tested individually (100 nM total RNP concentration) and in combination (100 nM total RNP concentration: 33 nM each of RNP 2, RNP 4, and RNP 21) against RNA isolated from human airway organoids (left), H1N1 influenza A (center), and influenza B (right). No target RNA RNP alone controls are denoted as “RNP 2,” “RNP 4,” “RNP 21,” and “RNP 2+4+21.” Background correction of fluorescence was performed by subtraction of reporter alone fluorescence values. Data are represented as mean ± standard error of the difference between means of three technical replicates. See also Figures S2A–S2D.
(C) RNA from five nasopharyngeal swabs confirmed negative for SARS-CoV-2 by RT-qPCR was tested against RNP 2+4+21 (100 nM total RNP concentration). The no target RNA RNP control is denoted as “RNP 2+4+21.” Raw fluorescence values over 2 h is shown. Data are represented as mean ± SD of three technical replicates. See also Figure S2E.
(D) Dilutions of genomic SARS-CoV-2 RNA independently quantified by BEI using ddPCR was tested against RNP 2+4+21 to determine the limit of detection (n = 20, technical replicates). Slope of the raw fluorescence curve over 2 h was calculated by performing simple linear regression of data merged from replicates and is shown as slope ± 95% confidence interval (left). Slopes were compared to the no target RNA RNP alone control using ANCOVA: ∗∗∗∗p < 0.0001. An individual reaction containing the diluted SARS-Cov-2 RNA was compared with the reaction without the target RNA and the number of true positives was counted at the 95% confidence level (right).
(E) Pre-extracted RNA from five nasopharyngeal swabs confirmed positive for SARS-CoV-2 by RT-qPCR was tested against RNP 2+4+21 (100 nM total RNP concentration) (n = 3, technical replicates). A confirmed negative swab was tested against RNP 2+4+21 for comparison. We added 0.3 μL of RNA from Patient Swabs 1–4, 0.26 mL of RNA from Patient Swab 5, and 0.3 μL of RNA from a confirmed negative swab to each 20 μL Cas13a reaction (in triplicate). Slope of the raw fluorescence curve over 2 h was calculated by performing simple linear regression of data merged from replicates and is shown as slope ± 95% confidence interval. Slopes were compared to the negative swab RNP background control using ANCOVA: ∗∗∗∗p < 0.0001. See also Figure S2F.
(F) The Ct value (average Ct count using CDC N1/N2 primers in RT-qPCR), the copies/mL of the original sample determined by qPCR, and the copies/μL in the Cas13a reaction are described for the RNA samples from each positive swab used in Figure 3E.