Figure S2.
Combining crRNAs improves SARS-CoV-2 detection, related to Figure 3
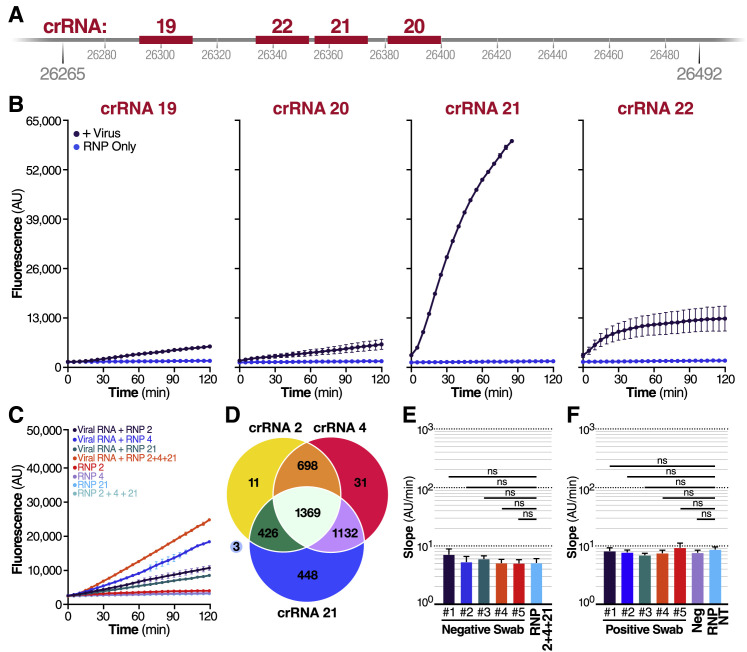
(A) Schematic of the SARS-CoV-2 envelope (E) gene, and the corresponding location of each crRNA spacer region.
(B) Cas13a RNPs made individually with each E gene crRNA (final RNP complex concentration of 100 nM) were tested against genomic SARS-CoV-2 RNA. Background fluorescence by the RNP alone in the absence of target RNA is shown as “RNP Only.” Raw fluorescence values over 2 h are shown. Data are represented as mean ± SD of three technical replicates.
(C) RNPs made with crRNA 2, crRNA 4, and crRNA 21 individually and in combination (100 nM total RNP concentration for each reaction) were tested against 1.5 × 104 copies/mL of extracted SARS-CoV-2 RNA, and compared to fluorescence from no target RNA RNP alone controls (“RNP 2,” “RNP 4,” “RNP 21,” and “RNP 2+4+21”). Raw fluorescence values over 2 h is shown. Data are represented as mean ± SD of three technical replicates.
(D) 4118 complete SARS-CoV-2 genome sequences deposited in NCBI RefSeq under taxonomy ID (2697049) were downloaded on 06/05/2020. Each crRNA was compared against the downloaded genomes for genomes with zero mismatches to each individual crRNA. The Venn diagram shows how many complete genomes have 100% homology to crRNAs 2, 4, and 21, as well as the overlap between crRNAs.
(E) Slope of the curves in Figure 3C over two h was calculated by performing simple linear regression of data from each replicate (n = 3) individually. The mean of the replicate slopes is shown as slope ± 95% confidence interval. Slopes were compared to the no target RNA RNP alone control using repeated-measures one-way analysis of variance (ANOVA) and Dunnett’s multiple comparisons test: ns = not significant.
(F) The same swabs as in Figure 3E were tested against a non-targeting crRNA (RNP NT) (final RNP complex concentration of 100 nM). Slope of the raw fluorescence curve over 2 h was calculated by performing simple linear regression of data merged from replicates (n = 3) and is shown as slope ± 95% confidence interval. Slopes were compared to the no target RNA RNP alone control using ANCOVA: ns = not significant.