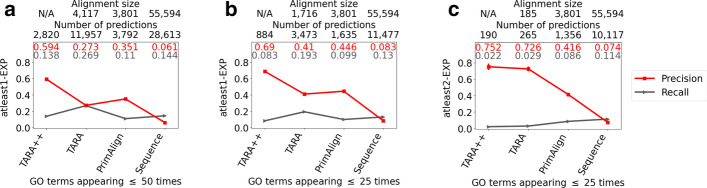
Fig. 5.
Comparison of TARA++ and three existing methods in the task of protein functional prediction. Comparison of TARA++ and three existing methods in the task of protein functional prediction, for rarity thresholds a 50 and b, c 25, and for ground truth datasets a, b atleast1-EXP and c atleast2-EXP. The alignment size (the number of aligned yeast-protein pairs) and number of functional predictions (predicted protein-GO term associations) are shown for each method, except that TARA++ does not have an alignment per se. i.e., TARA++ comes from the overlap of predictions made by TARA and TARA-TS; hence the “N/A”s. For TARA++ and TARA, results are averages over all balanced datasets; the standard deviations are small and thus invisible. Results for the other ground truth-rarity datasets are shown in Additional file 1: Fig. S11