Fig. 1.
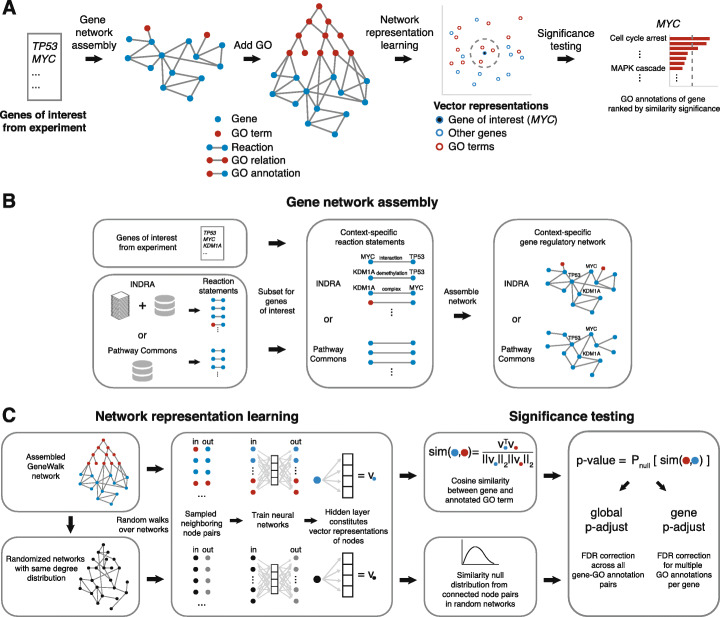
GeneWalk methodology. a Schematic introducing the key aspects of the GeneWalk method. The input is a list with genes of interest, e.g., all differentially expressed genes under a certain experimental condition. Using the INDRA [21, 37] or Pathway Commons [18, 43] knowledge base, all molecular reactions in which these genes are involved are retrieved and assembled in a condition-specific gene regulatory network, to which GO ontology and annotations are then connected. Through network representation learning, each gene and GO term can be represented as vectors, suitable for similarity significance testing. For each gene, GeneWalk gives as output the similarities with all of the connected GO terms, along with their significance, specifying which annotated functions are relevant under the experimental condition. b Schematic details of the gene network assembly procedure from the input list with genes of interest and knowledge base INDRA or Pathway Commons (PC). These knowledge bases provide reaction statements. INDRA accumulates these from automated literature reading and database queries [21, 37], while Pathway Common only queries databases [18, 43]. Another difference between INDRA and PC is that INDRA also provides gene–GO relations through automated text mining. Then a strict subset results in the collection of context-specific reaction statements that involve only genes of interest. These reaction statements are then assembled into a gene regulatory network. c Schematic details of the network representation learning and significance testing parts of GeneWalk. Random walks are generated from the assembled GeneWalk Network (GWN), yielding a large collection of sampled neighboring node pairs, which form the training set of (input,output) of a fully connected shallow neural network (NN) where each node from the GWN is represented as a single feature. The learned hidden layer is the vector representation of a node. And the similarity of a node pair then equals the cosine similarity between the corresponding node vectors. To enable similarity significance testing, we generated randomized networks that were also subjected to DeepWalk and whose resulting cosine similarity values form the null distributions used to calculate a p value of the experimental similarities between a gene and GO term node. Finally, because multiple GO terms were tested, we applied two FDR corrections that address different questions. The gene p-adjust values rank the context-specific relevance of all annotated GO terms for a pre-defined gene of interest. The global p-adjust can be used to identify relevant genes and their functions across the whole input gene list