FIGURE 2.
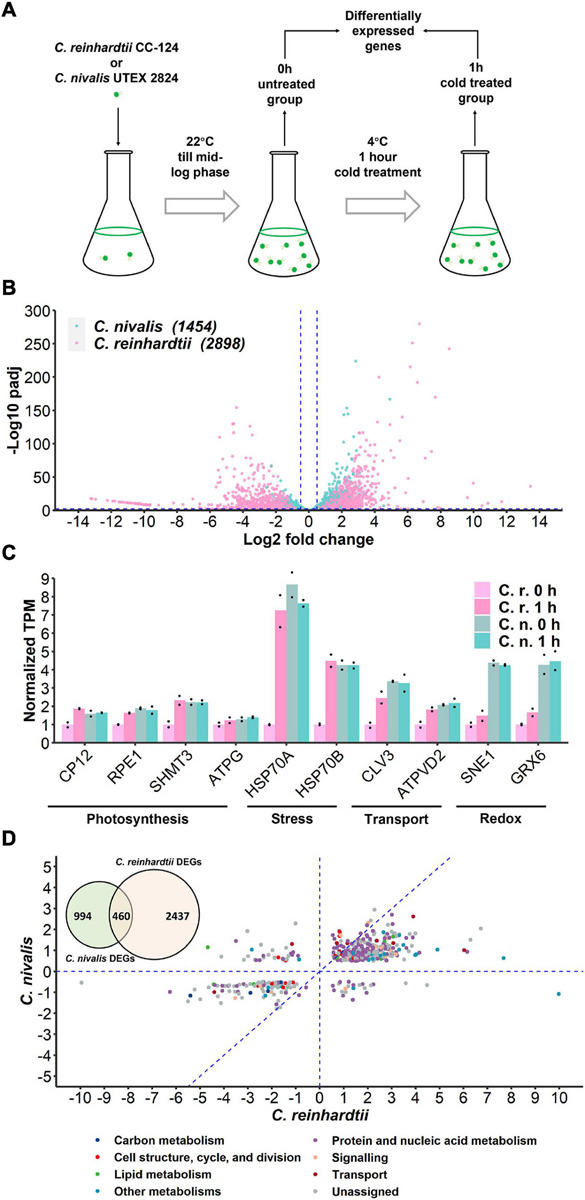
Transcriptome of C. nivalis is relatively stable compared to C. reinhardtii. (A) Schematic of sampling for RNA-seq. Algal cells were grown at 22°C until mid-log phase (0 h) and transferred to 4°C immediately for 1 h. Samples at 0 h and 1 h were collected for RNA-seq and differential gene expression analysis. (B) Volcano plot of DEGs in C. nivalis and C. reinhardtii. The blue dashed lines indicate the threshold (fold change > 20.5, adjusted P-value < 0.01) for DEGs filtering. The number of DEGs is indicated in parentheses after the species name. padj, FDR-adjusted P-value. (C) Transcripts per million reads (TPM) of ten conserved genes. TPM values were normalized to TPM value of C. reinhardtii at 22°C. The dots indicate the two replicates for each bar. (D) Venn diagram in the upper-left corner shows overlapping DEGs in C. nivalis and C. reinhardtii. Scatter plot shows fold changes of the overlapping DEGs in the two species separately and the category of DEG is indicated by color. The blue dash lines (x = 0, y = 0, x = y) partition the DEGs to clarify the relationship between fold changes in C. nivalis and C. reinhardtii. LFC, log2 fold change.
