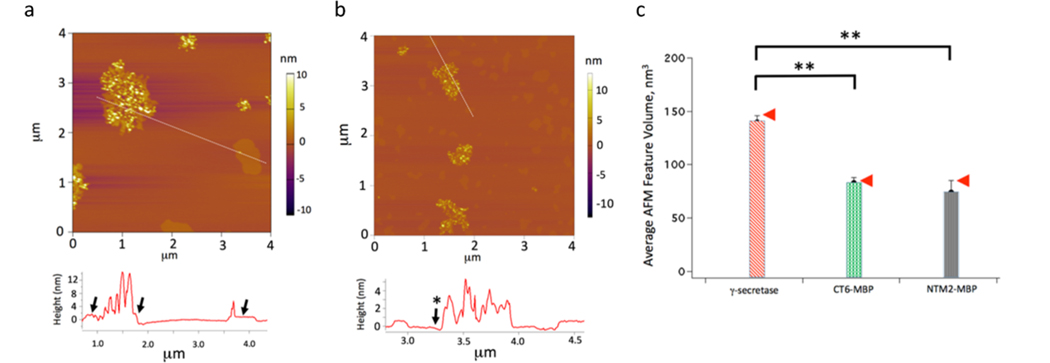
Figure 4.
APP and Notch substrates locate in different domains of the lipid bilayer. (a) Insertion of MBP-SB4 into the model membrane; the image was taken at 31 °C after the addition of the substrate for 150 min. The image line profile is obtained from the indicated line segment. (b) Insertion of MBP-NTM2 into the model membrane; the image was taken at 31 °C after the addition of the substrate for 150 min. (c) Bar chart obtained from computing the averae AFM feature volume for γ-secretase (n = 86), MBP-SB4 (n = 50), and MBP-NTM2 (n = 50) for the subset of features consistent with arising from single molecules protruding from the supported membrane. The arrows indicate, for reference, the predicted volume of protruding γ-secretase subunit nicastrin (NCT; 147 nm3/molecule) and MBP (85 nm3/molecule) based on polymer size arguments (Figure S6, the Supporting Information section). The double asterisks in Figure 4c indicate statistically significant differences between the average feature volumes (p < 0.05).