Figure 4.
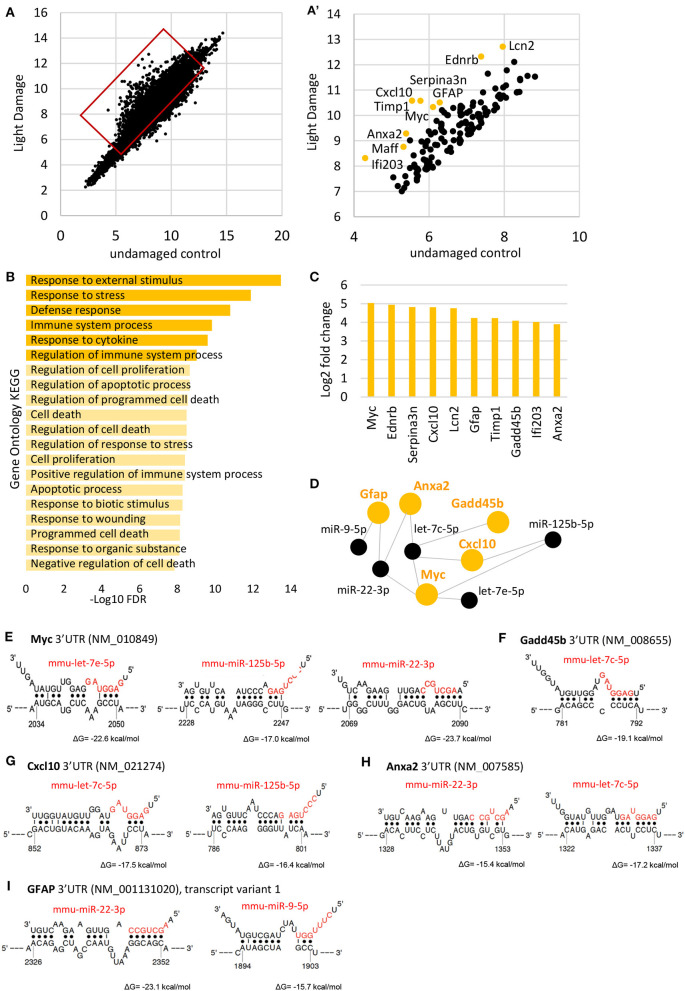
Müller glia gene expression after light damage. (A) Scatterplot of genes expressed in undamaged controls and 7 days after light damage (LD) from microarray (log2 of relative expression). (A′) Scatter plot of the top 100 highly expressed genes after light damage. (B) Gene ontology of the top 100 genes upregulated after light damage using ShinyGO v0.61. (C) Expression levels of the top 10 genes upregulated after light damage compared to undamaged controls (five and six biological replicates, respectively, one technical replicate per condition). (D) DIANA-TarBase v.8 analysis of the top genes upregulated after light damage and MG miRNAs that declined and are likely to target these genes based on prediction and experimental confirmation. (E–G) Binding sites of identified miRNAs from D (DIANA-TarBase) in the 3′UTRs of Myc (E), Gadd45b (F), Cxcl10 (G), Anxa2 (H), and Gfap (I) using STarMirDB. The seed sequences of the particular miRNA are shown in red. Dots represent complimentary base pairing. ΔG indicates the structural accessibility at the target site, low values (e.g., −17 kcal/mol) indicate high accessibility.