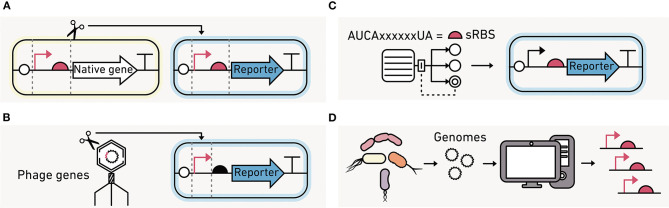
Figure 11.
Strategies for identifying regulatory elements to program environmental microbes. (A) Native promoters and RBS from the organisms being engineered can be used to build the synthetic circuits if their activities are known. (B) Promoter and polymerase pairs that are functionally orthogonal from those in the native organisms can be used to control transcription, such as those encoded by phage. (C) Computational models can design synthetic RBS sequences (sRBS) with a range of translation initiation strengths. (D) Large-scale transcriptomics data can be mined for promoter and RBS sequences that are active at similar levels across a broad range of strains.