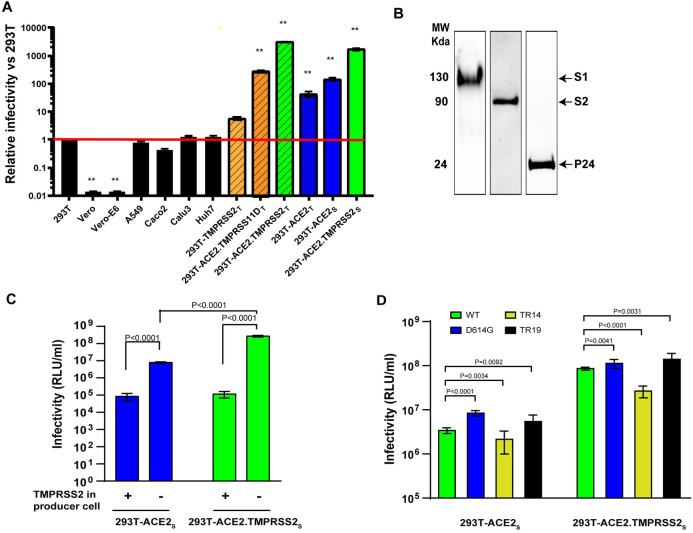
Fig 1. SARS-CoV-2 lentiviral pseudovirus infectivity under various conditions.
(A) Relative infectivity of SARS-CoV-2 pseudoviruses in various target cells. The Y-axis shows relative infectivity compared to background in 293T cells. Subscript ‘t’ or ‘s’ refers to transient or stable expression, respectively. Red line indicates background level (1 x 104 relative units of luciferase activity/milliliter or RLU/ml). (B) Western blots probed for spike S1/S2 subunits and HIV p24 incorporated into pseudoviruses. Lanes represent different blots (C) Infectivity on 293T-ACE2s and 293T-ACE2.TMPRSS2s of pseudoviruses primed with or without TMPRSS2 during pseudovirus production. (D) Infectivity on 293T-ACE2s and 293T-ACE2.TMPRSS2s of pseudoviruses bearing full-length, wildtype S glycoprotein (WT), an S glycoprotein with the D614G substitution, or S glycoproteins with C-terminal truncations of 14 (TR14) and 19 (TR19) amino acids. Data are shown as means and standard deviations from 3–4 independent experiments (panel A, C and D). The tests for two-group comparison were analyzed using GraphPad Prism software. P values of 0.05 were considered statistically significant. **: P<0.0001, compared to the infectivity in 293T.