Figure 3.
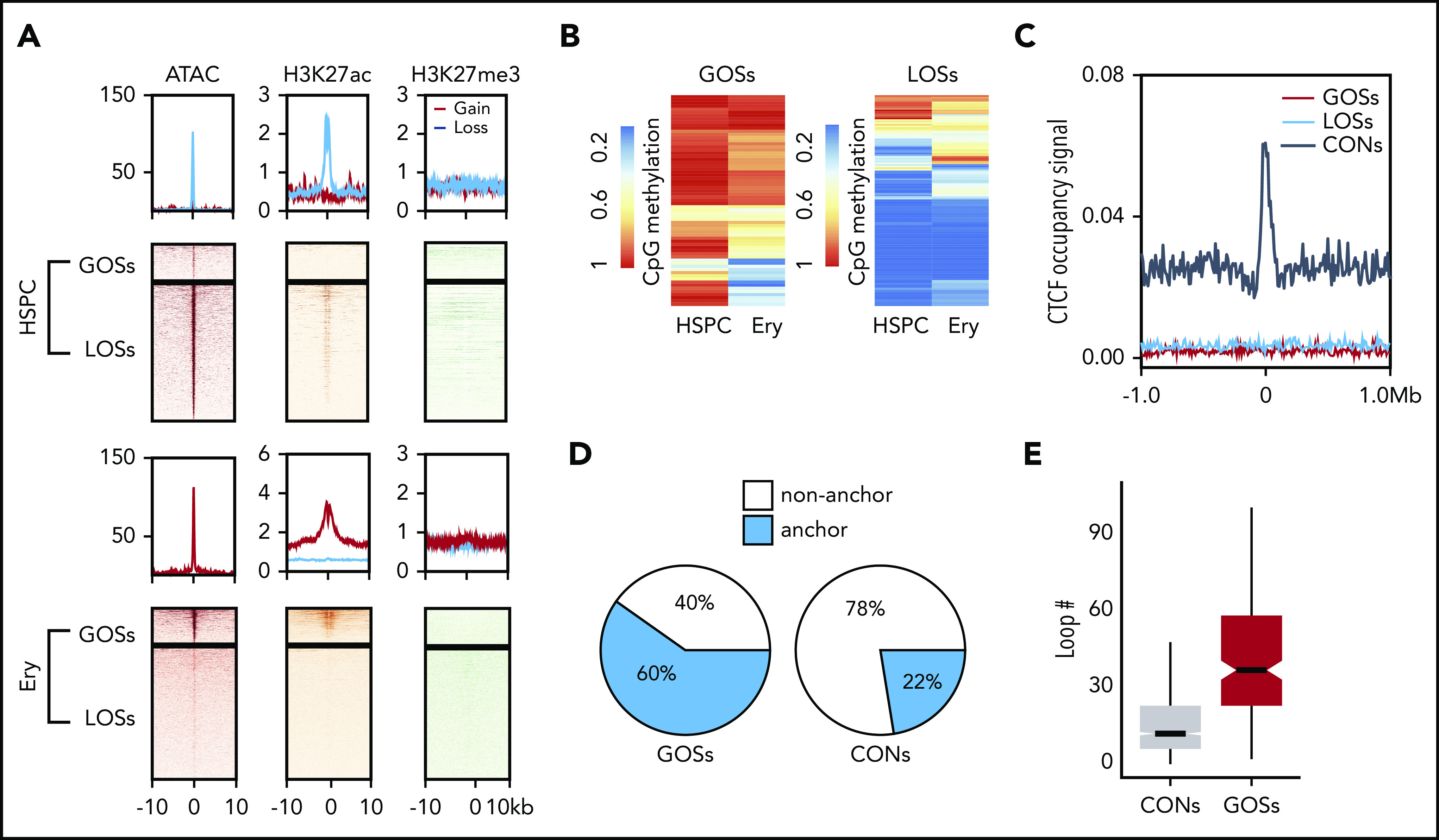
Epigenetic and 3D chromatin features of dynamic CTCF occupancy. (A) Heatmap showing the signals of ATAC-seq, H3K27ac, and H3K27me3 around the 20-kb windows centered on the peak summits of GOSs (top) and LOSs (bottom) in HSPCs and erythroblasts. (B) Heatmap showing the methylation levels of CpG sites within the CTCF motifs in HSPC and erythroblast dynamic sites. Each row represents 1 CpG site. Left panel represents the CpG sites in GOSs. Right panel represents the CpG sites in LOSs. (C) Aggregated distribution of different groups of CTCF-binding sites within a 2-Mb window of the TAD boundary in erythroblasts identified by Hi-C. (D) Pie chart showing the proportion of CTCF peaks overlapping with the anchors of chromatin loops identified by H3K27ac HiChIP using HSPC differentiated erythroblasts from the same donor. (E) Box plot showing the distribution of the number of loops per anchor region for GOSs and CONs in erythroblasts.
