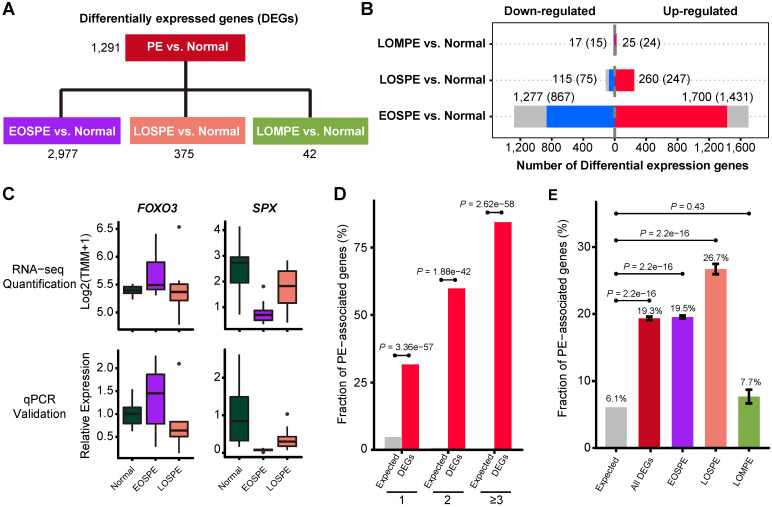
Figure 2.
Differentially expressed genes in preeclamptic placentae. (A) The differentially expressed genes (DEGs) for each group were identified using two methods, DESeq2 and edgeR. The 4 comparisons of the RNA-seq data include: all PE samples vs. normal controls, EOSPE samples vs. controls, LOSPE samples vs. controls, or LOMPE samples vs. controls. (B) The red bars represent up-regulated genes, the blue bars represent down-regulated genes and the grey bars non-coding genes. Numbers represent DEGs in each group, and numbers in brackets represent protein-coding DEGs. (C) DEGs verified by qPCR. FOXO3: RNA-seq quantification boxplot at top-left panel and qPCR validation result at bottom-left panel. SPX: RNA-seq quantification boxplot at top-right panel and qPCR validation result at bottom-right panel. (D) The enrichment of PE associated genes in DEGs. PE-associated genes were curated from literature (Table S4). Of the genes that appear once, twice and three or more times in literature, 31.7%, 59.9% and 84.4% were recovered by the DEGs we identified in PE. The X-axis represents the number of times of a gene reported in the literature, the Y-axis represents the fraction of reported PE-associated genes under each category, with red bars representing fractions of PE-associated genes that appear once, twice, and three or more times in different literature, and gray bars the expected fractions. (E) The enrichment of known PE-associated genes in protein-coding DEGs in EOSPE (2,298 genes), LOSPE (322 genes), LOMPE (39 genes) and all DEGs (2,405 genes). We collected 1,177 PE-associated genes from the literature (Table S4 and Methods), and calculated the their enrichment in DEGs of different groups: red bar represents the fraction of PE-associated genes in all DEGs (19.3%, P = 2.2e-16), purple bar EOSPE (19.5%, P = 2.2e-16), pink bar LOSPE (26.7%, P = 2.2e-16) and light-green bar LOMPE (7.7%, P = 0.4261), compared to gray bar which represents the expected ratio (6.1%) of 1,177 PE-associated genes to 19,351 human coding genes (GRCH38.p12 ENSEMBL Gene V93). One-sided Fisher's exact test was used to compare the nodes kept with the nodes removed. Error bars represent the standard error of the fraction, estimated using bootstrapping method with 100 resamplings.