Figure 5.
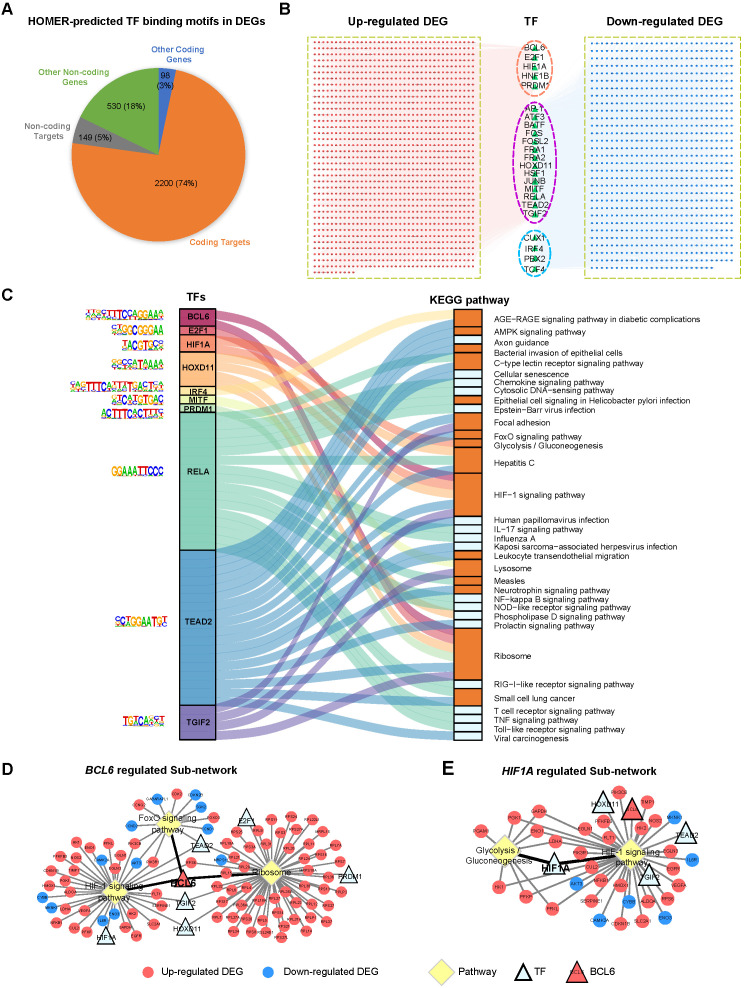
Identification of critical transcription factors that contribute to the dysregulation of pathways involved in EOSPE. (A) Transcription factor-binding motifs were searched in the DEGs of EOSPE using HOMER. Of 2977 DEGs in EOSPE, 79% (2349/2977) were predicted to be targeted by 23 TFs. (B) TF-targets network for the DEGs of EOPSE. Of the 23 enriched TFs (the triangles), 5 TFs (pink circles) target the up-regulated DEGs, 4 TFs (blue circles) target the down-regulated DEGs, and 14 TFs (purple circles) target the both up- and down-regulated DEGs. The up-regulated targets are in the red nodes (1454), and the down-regulated targets are in the blue nodes (895). (C) The Sankey diagram showing the relationship between TFs and the enriched pathways with its targets. Enrichment analysis was performed on the targets of each TF. A total number of 34 enriched pathways (the right column) were found for 10 TFs (colored column in the left). The binding motifs corresponding to TFs are listed on the left. The orange color in the right bar indicates the pathways that are overlapped with those enriched with DEGs of EOSPE (Figure 3A), and the light blue color in the right bar indicates the pathways that are newly found enriched pathways with the targets of TFs. (D-E) 'TF-Target-Pathway bipartite networks' for TFs BCL6 and HIF1A. Most of BCL6 targeting DEGs are involved in three KEGG pathways (D). The HIF1A targeting DEGs are involved in two pathways (E). Circles: non-TF genes; triangles: TFs; yellow diamonds: KEGG pathways; red: up-regulation; blue: down-regulation; light blue: no expression change.