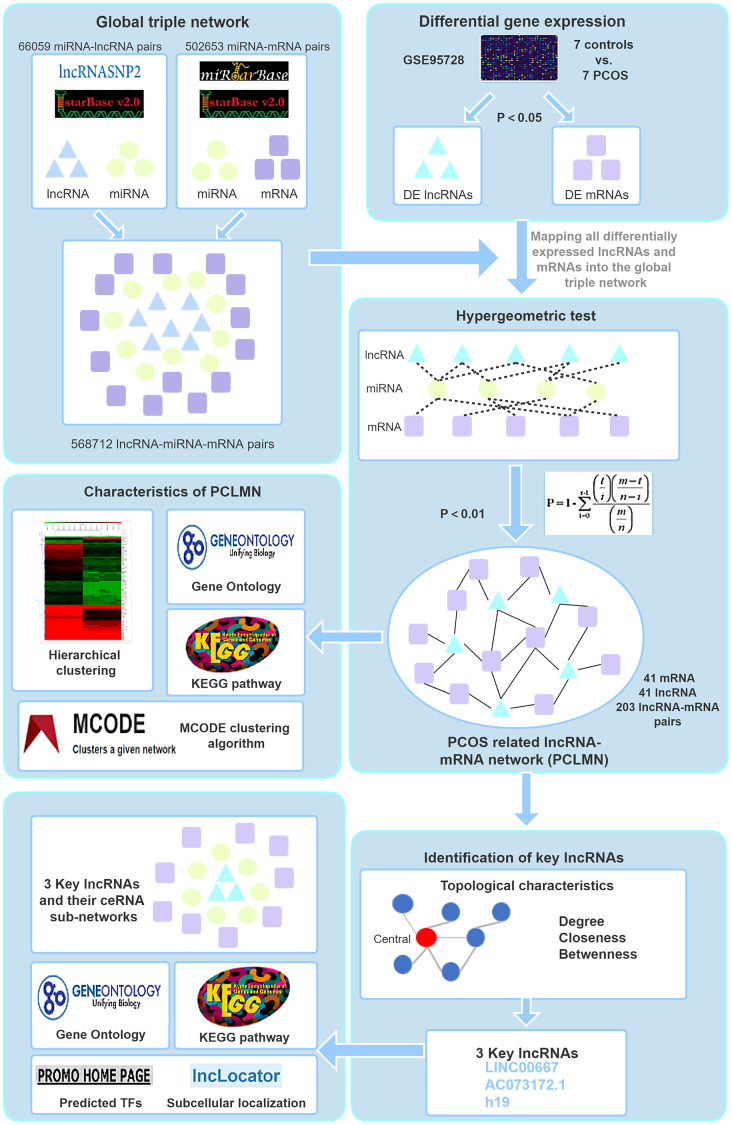
Figure 2.
Study workflow. First, we constructed a global background network based on predicted lncRNA-miRNA and miRNA-mRNA pairs. Second, we applied a hypergeometric test to construct the PCLMN and performed network topology analysis to determine the lncRNAs with the highest centroid variability. Lastly, we explored the subcellular localization of the key lncRNAs thus identified, performed functional module analyses, and identified putative transcription factors regulating the expression of the candidate lncRNAs.