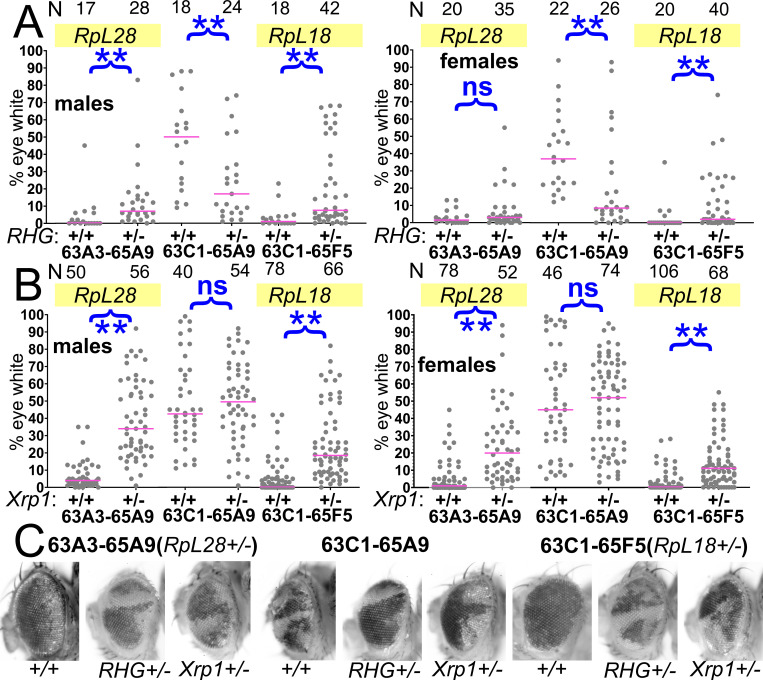
Figure 5. Contributions of apoptosis and Xrp1 to elimination of segmentally aneuploid cells.
Panels A-B show the contribution of w- segmentally aneuploid cells to adult eyes of indicated genotypes, N as indicated for each genotype. (A) Heterozygosity for Df(3L)H99, which deletes the proapoptotic genes rpr, hid, and grim (‘RHG’) significantly increases contribution of Df63A3-65A9/+ and Df63C1-65F5/+ cells to adult eyes. These genotypes approach the contribution level of Df63C1-65A9/Df(3L)H99 cells which do not affect any Rp locus the results for 63A3-65A9/Df(3L)H99, 63C1-65A9/Df(3L)H99, and 63C1-65F5/Df(3L)H99 cannot be distinguished in males: KruskalWallis test, p=0.18; in females, this hypothesis is rejected and post-hoc testing identified the 63C1-65A9/Df(3L)H99 data as different from the other genotypes (p=0.018 for the comparison to 63A3-65A9/H99, p=0.027 for the comparison to 63C1-65F5/H99). (B) Heterozygosity for Xrp1, which is required for the slow growth and cell competition of Rp+/-point-mutant cells, significantly increases contribution of Df63A3-65A9/+ and Df63C1-65F5/+ cells to adult eyes. Xrp1 did not affect the contribution of Df63C1-65A9/+ cells that do not affect any Rp locus. We also tested the hypothesis that Df(3L)H99, Xrp1 mutation, and an RpS3/+ genetic background (see Figure 8) suppressed cell competition to an equal degree. This hypothesis was rejected for both males and females of Df63A3-65A9/+ and Df(3L)63C1-65F5/+ (Kruskal Wallis test p=2.9×10−17, 9.7 × 10−21, 2.1 × 10−5, and 1.8 × 10−7, respectively). For each deletion, all the genotypes were individually significantly different except Df(3L)63C1-65F5/H99 and Df(3L)63C1-65F5/Xrp1 males, which were not significantly different (p=0.06, Conover post-hoc test with Holm correction). (C) Representative examples of these genotypes. Statistics. Pairwise comparisons using the Mann-Whitney procedure with the Benjamini-Hochberg correction for multiple testing (see Supplementary file 2). ns – difference not statistically significant (p>0.05). ** - difference highly significant (p<0.01). Multiple comparisons using the Kruskal Wallis test as described for panel B. Genotypes. For 63A3-65A9 in panel A: y w hsF; PBac{WH}f01922 P{XP}d02570 /FRT80B and y w hsF; PBac{WH}f01922 P{XP}d02570 /Df(3L)H99 FRT80B. For 63C1-65A9 in panel A: y w hsF; PBac{WH}f05041 P{XP}d02570 /FRT80B and y w hsF; PBac{WH}f05041 P{XP}d02570 /Df(3L)H99 FRT80B. For 63C1-65F5 in panel A: y w hsF; PBac{WH}f05041 P{XP}d02813 /FRT80B and y w hsF; PBac{WH}f05041 P{XP}d02813 /Df(3L)H99 FRT80B. For 63A3-65A9 in panel B: y w hsF; PBac{WH}f01922 P{XP}d02570 /FRT82B and y w hsF; PBac{WH}f01922 P{XP}d02570 /FRT82B Xrp1m2-73. For 63C1-65A9 in panel B: y w hsF; PBac{WH}f05041 P{XP}d02570 /FRT82B and y w hsF; PBac{WH}f05041 P{XP}d02570 /FRT82B Xrp1m2-73. For 63C1-65F5 in panel B: y w hsF; PBac{WH}f05041 P{XP}d02813 /FRT82B and y w hsF; PBac{WH}f05041 P{XP}d02813 /FRT82B Xrp1m2-73.