Fig. 1.
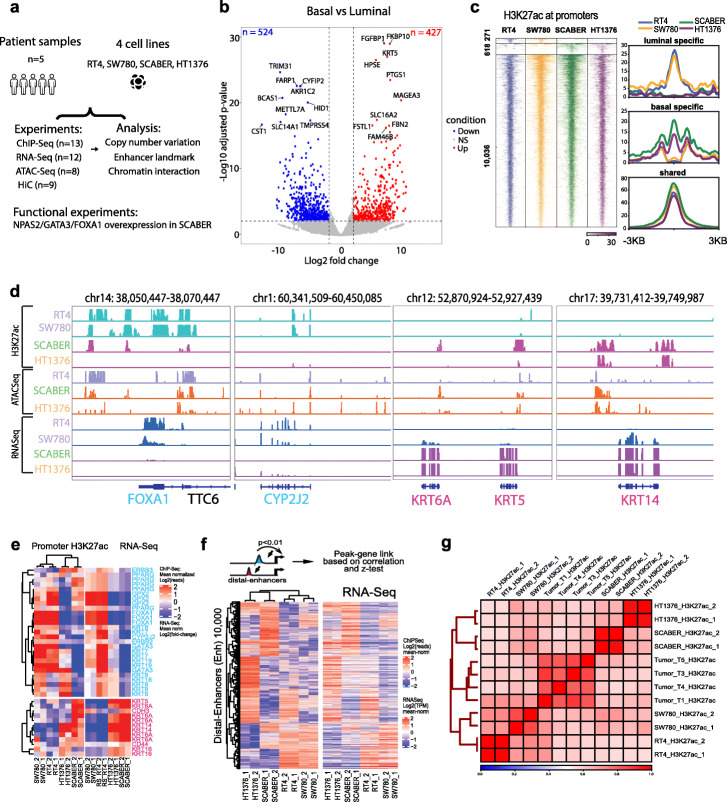
Luminal and basal transcriptional BLCA subtypes are associated with distinct promoter and distal enhancers’ activity at the epigenetic level. a Overall design of the study. b Differential expression gene (DEG) analysis of luminal cell lines (RT4 and SW780) and basal cell lines (SCABER and HT1376) shows 427 basal-specific upregulated genes and 524 luminal-specific upregulated genes. c Heatmap of differential H3K27ac ChIP-Seq at promoters (left). Signal H3K27ac intensity profiles for each cluster of BLCA cells (right). d Genome browser signal tracks for a panel of luminal and basal genes. Shown here are the tracks of H3K27ac ChIP-Seq, ATAC-Seq, and RNA-Seq data in RT4, SW780, SCABER, and HT1376 cells. e Promoter H3K27ac and its associated RNA-Seq signals for selected luminal and basal genes shows remarkable similarity. f Integrated H3K27ac peaks at distal enhancers and RNA-Seq gene expression association model identifies putative enhancers and gene regulation. Top 10,000 most variable enhancers (left heatmap) are plotted along with their corresponding gene expression (right heatmap). g Correlations of genome-wide H3K27ac signals between the bladder cancer cell lines and tumor samples demonstrate similarity of enhancer landscape