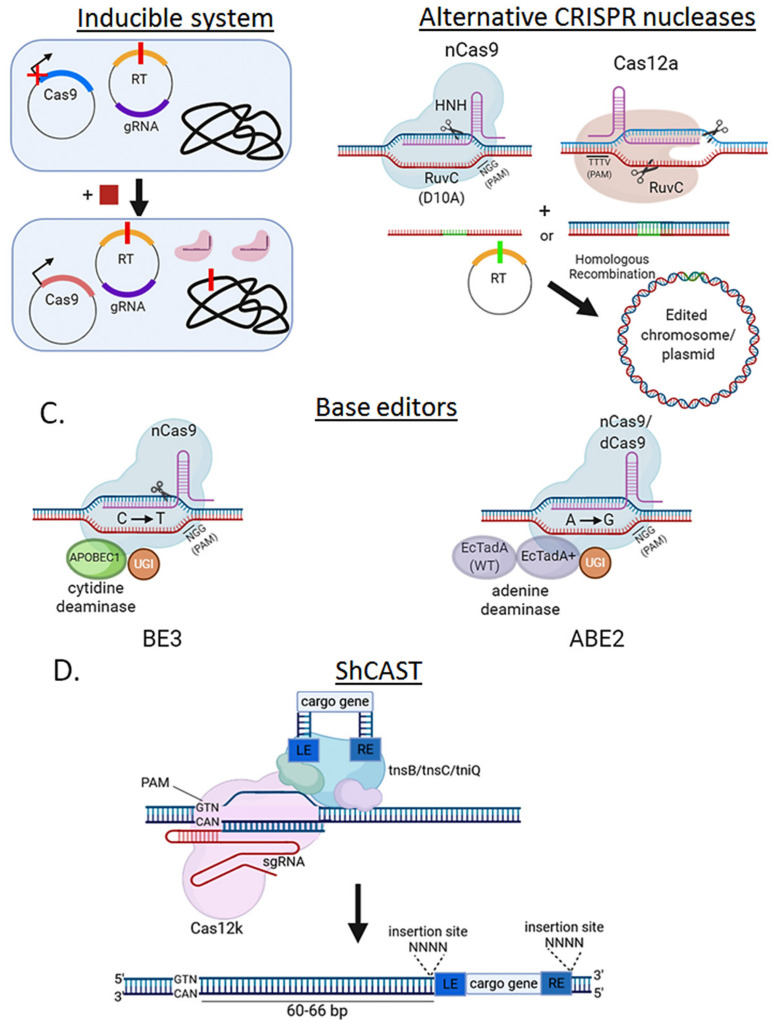
Figure 5.
Alternative strategies to circumvent SpCas9 cytotoxicity. (A) Use of inducible systems to express SpCas9. Via an inducible promoter, SpCas9 expression is strongly repressed without inducer present (square) and only induced after exponential culture so that enough cells can survive and perform the genome edit. (B) Using less toxic nucleases to achieve editing. nCas9, which only cleaves one strand of DNA, and Cas12a (PAM: TTTV, where V is A or C or G) can be less toxic than SpCas9. (C) SpCas9-derived base editors eliminate the double-stranded break requirement for genome editing. A translational fusion of nCas9 (nickase) or dCas9 (“dead”), a cytidine (e.g., APOBEC1 in BE3) or adenosine (e.g., TadA-EcTadA+ in ABE2) deaminase domain, and an uracil DNA glycosylase inhibitor (UGI) is introduced on a plasmid into the cell. Upon nuclease binding and DNA strand unwinding, cytidines (or adenines) on the non-target strand within a defined window adjacent to the PAM are rapidly converted to uracil (or inosines), which is then processed as thymidine (or guanines) by DNA polymerase. (D) ShCAST insertion mechanism. A Tn7-like transposon from Scytonema hoffmani encodes transposases (tnsB, tnsC, tniQ), a nuclease deficient type V CRISPR protein (Cas12k) and guide RNA. This complex is combined with a cargo gene flanked by LE and RE elements. ShCAST is directed to the target locus and integrates the cargo gene 60–66 bp downstream of the PAM sequence, generating and insertion of the cargo gene flanked by the SE and RE elements, and a duplicated (4 bp) insertion site.