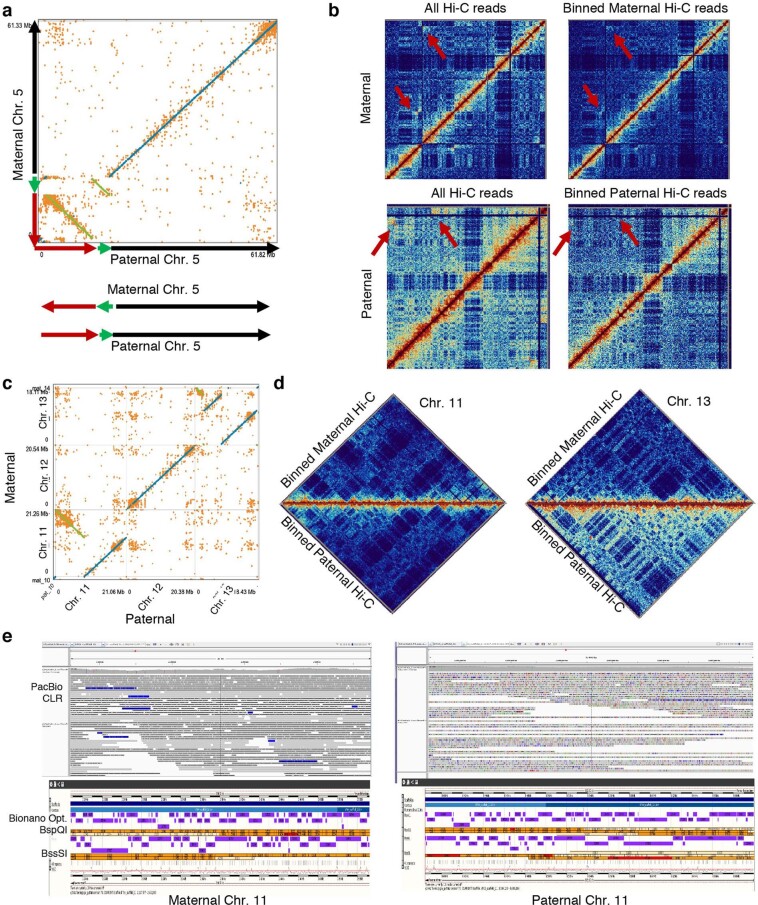
Extended Data Fig. 10. Large haplotype inversions with direct evidence in the zebra finch trio assembly.
a, Two inversions (green and red) in chromosome 5 found from the MUMmer NUCmer107 alignment of the maternal and paternal haplotype assemblies, visualized with dot108. b, Hi-C interaction plot showing that the trio-binned Hi-C data remove most of the interactions from the other haplotype (red arrows), which could be erroneously classified as a mis-assembly if only one haplotype was used as a reference. c, An 8.5-Mb inversion found on chromosome 11 and a complicated 8.1-Mb rearrangement on chromosome 13 between maternal and paternal haplotypes. d, No mis-assembly signals were detected from the binned Hi-C interaction plots, indicating that the haplotype-specific inversions are real. e, Half the PacBio CLR span and Bionano optical maps agree with the inversion breakpoints in chromosome 11, supporting the haplotype-specific inversion.