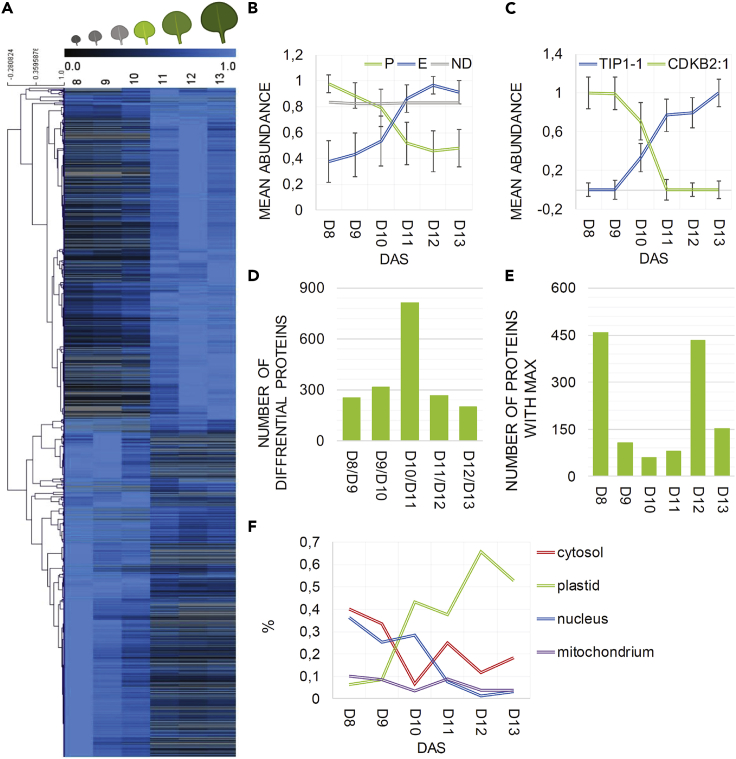
Figure 2.
Protein changes across early leaf development
(A) Heatmap of differential proteins identified using one-way ANOVA and absence/presence criteria (see results). Data were normalized to the maximal intensity. DAS, days after stratification. ANOVA analysis and heatmap were prepared using MeV software.10 Leaf panel was created using BioRender.com.
(B) Mean ± SD of all proteins classified as proliferation or expansion specific (see heatmap), in comparison with the mean of all non-differential (ND) proteins across leaf developmental time series (day 8 to day 13).
(C) Maximum normalized abundance of a selected proliferation and expansion marker proteins.
(D) Number of differential proteins calculated between two consecutive time points (e.g., between day 8 and day 9) using t test and absence/presence criteria (see results).
(E) Number of proteins with the highest abundance measured at a given day (x axis).
(F) SUBA database was used to assign subcellular localization.11 Percentage of proteins (where 100% is equal to 1) characterized by a given subcellular localization in comparison to all proteins peaking at a given time point (see Figure 1E).
In (A–F), data are from four replicas. In (D) and (E), analysis was restricted to the 1,215 proteins assigned as either proliferation or expansion specific using one-way ANOVA and absence/presence criteria. Complete data can be found in Data S1.