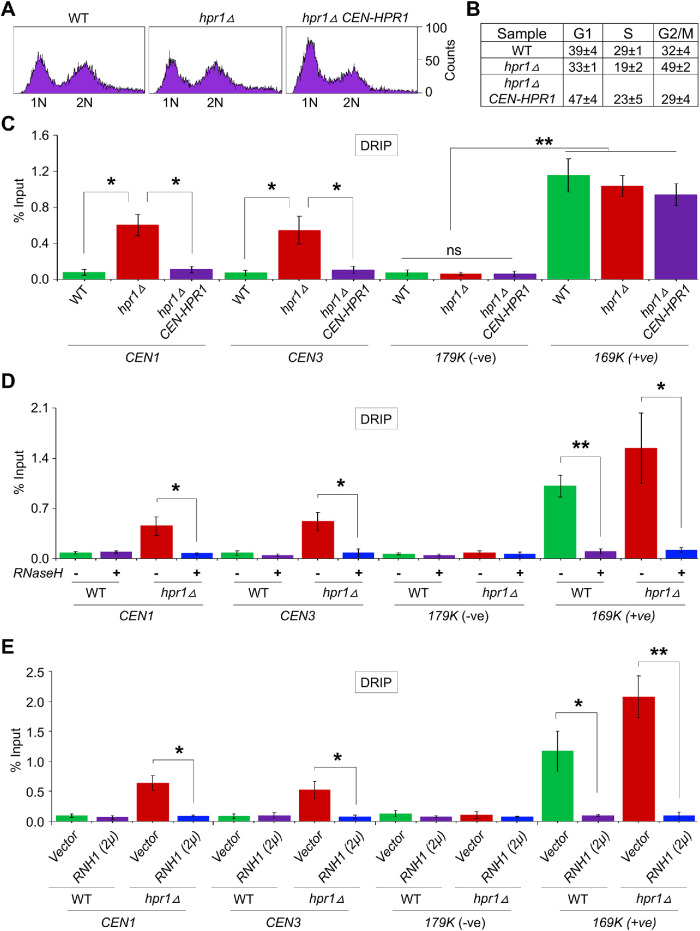
FIGURE 2:
Hpr1 prevents the accumulation of R-loops at the CEN chromatin. (A) FACS profile of logarithmically growing cultures of WT (YMB11246), hpr1Δ (YMB11247), and hpr1Δ CEN-HPR1 (YMB11248) cells used for DRIP experiments. (B) Cell cycle categorization of strains from A representing the percentage of cells in G1, S, and G2/M. (C) Accumulation of R-loops at CEN chromatin in hpr1Δ and its suppression by CEN-HPR1. DRIP analysis of yeast strains from A using the S9.6 (anti–DNA-RNA hybrid) antibody. Levels of R-loops (% input) at CENs (CEN1 and CEN3), a negative control (179K), and a positive control (169K) were determined by DRIP-qPCR. Average from three biological replicates ± SE. **p value < 0.01, *p value < 0.05, ns = statistically not significant, Student’s t test. (D) Accumulation of R-loops at CEN chromatin in hpr1Δ cells with (+) or without (–) treatment with RNase H. DRIP analysis of WT (JG595) and hpr1Δ (YMB11097) using the S9.6 (anti–DNA-RNA hybrid) antibody. Levels of R-loops and statistical significance were determined as described in C. (E) Accumulation of R-loops at CEN chromatin in hpr1Δ cells with or without RNH1 overexpression. DRIP analysis using the S9.6 (anti–DNA-RNA hybrid) antibody was performed for a WT strain with vector (YMB11210) or 2μ-RNH1 (YMB11211) and an hpr1Δ strain with vector (YMB11212) or 2μ-RNH1 (YMB11213). Levels of R-loops and statistical significance as described in C.