FIGURE 3:
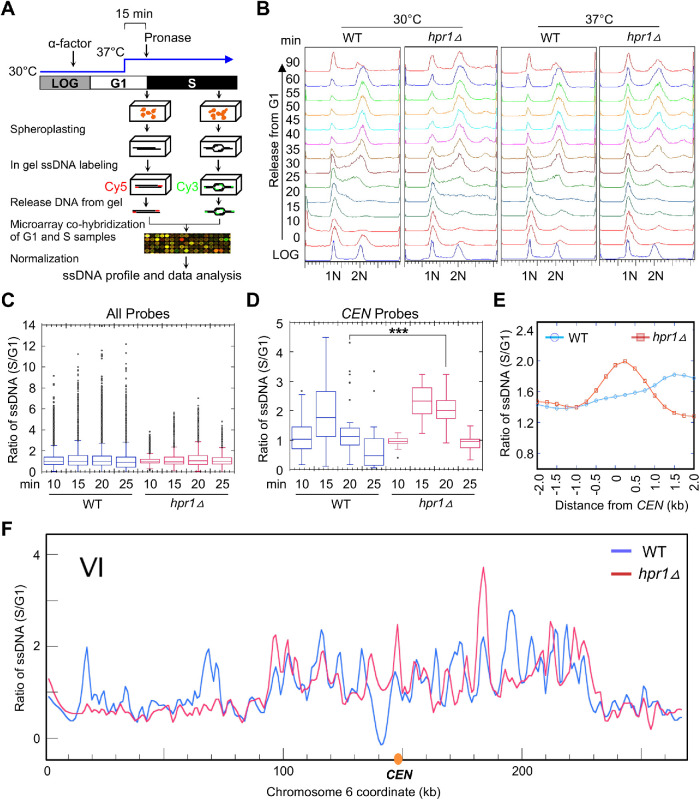
Genome-wide ssDNA mapping shows accumulation of ssDNA at CENs in the hpr1Δ strain at the restrictive temperature (37°C). (A) Schematic presentation of genomic ssDNA mapping. Details of cell culture conditions, ssDNA labeling, and quantification are described in Materials and Methods. Briefly, cells synchronized in G1 by α-factor at 30°C were acclimatized to 37°C for 15 min before the release into S-phase by Pronase treatment. S-phase samples were collected every 5 min starting at 10 min postrelease, as shown in B. Select S-phase samples (10, 15, 20, and 25 min post–G1 release) were subject to ssDNA labeling, and each was cohybridized with a ssDNA labeled G1 control sample onto the microarray. (B) FACS profile of WT (BY4741) and hpr1Δ (BY4741 hpr1Δ) strains. Cells were collected as described in A. The positions of G1 and G2 cells are indicated by “1N” and “2N”, respectively. (C, D) Box plots for quantification of ssDNA from cells sampled at the indicated time points post–release from G1- into S-phase for all probes on the microarray (C) and for probes overlapping only with the CEN (D). Statistical analysis was performed by analysis of variance followed by Tukey’s multiple comparison. ***p value < 0.001. (E) Meta-analysis of ssDNA in a 4 kb window centering on the CEN for cells sampled at 20 min post–release from G1- into S-phase. (F) Example of a chromosomal plot of ssDNA on chromosome VI from cells sampled at 20 min at 37°C. For all chromosome plots, see Supplemental Figure S2.