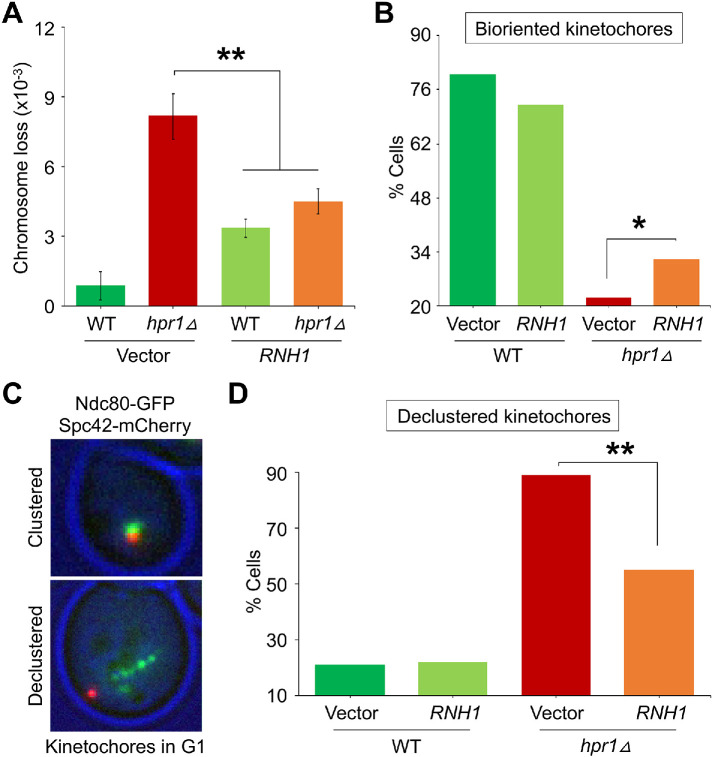
FIGURE 7:
Overexpression of RNH1 suppresses CIN, kinetochore biorientation, and kinetochore clustering defects in hpr1∆ strains. (A) RNH1 overexpression suppresses the frequency of chromosome segregation errors in the hpr1Δ strain. Frequencies of CF loss in WT with vector (YMB11206) or 2μ-RNH1 (YMB11207) and hpr1Δ with vector (YMB11208) or 2μ-RNH1 (YMB11209) strains were measured by a colony color assay as detailed in Materials and Methods. Average from three biological experiments ± SE. **p value < 0.01, Student’s t test. (B) RNH1 overexpression suppresses kinetochore biorientation defects in metaphase cells of hpr1∆ strains. WT with vector (YMB11494; n = 135 cells) or 2μ-RNH1 (YMB11495; n = 123 cells) and hpr1∆ with vector (YMB11496; n = 134 cells) or 2μ-RNH1 (YMB11497; n = 216 cells) strains containing Ndc80-GFP and Spc42-mCherry were grown at room temperature (25°C). The cutoff for metaphase spindle length was 1.7 μm. Spc42-mCherry was used as a spindle pole marker. Percentages of cells showing bioriented kinetochores are shown. Statistical significance was determined by χ2 test. *p value < 0.05. (C) Representative images of G1 cells depicting clustered and declustered kinetochores based on the position of Ndc80-GFP (green) and Spc42-mCherry (red). (D) RNH1 overexpression suppresses kinetochore clustering defects of hpr1∆ cells in G1. WT with vector (YMB11494; n = 104 cells) or 2μ-RNH1 (YMB11495; n = 103 cells) and hpr1∆ with vector (YMB11496; n = 100 cells) or 2μ-RNH1 (YMB11497; n = 292 cells) carrying Ndc80-GFP and Spc42-mCherry were grown at room temperature (25°C) to LOG phase, and G1 cells were selected based on the cell morphology (single-celled, no bud). Percentage of cells showing declustered kinetochores in G1 are shown. Statistical significance was determined by χ2 test. **p value < 0.01.