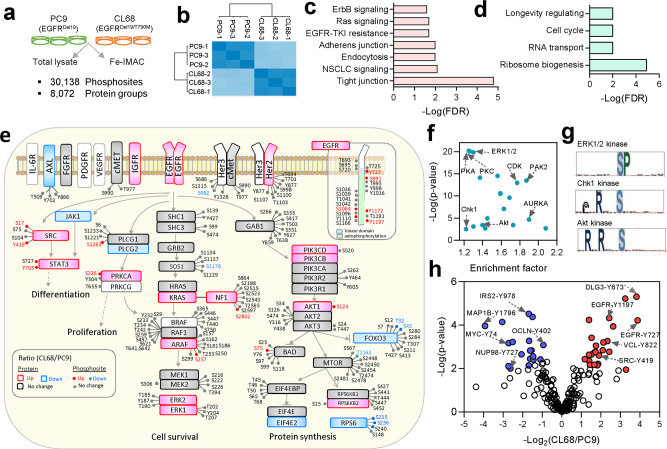
Fig. 5. Differential phosphoproteome profiling of EGFR-TKI-sensitive and resistant lung cancer cells.
a Experimental design and summary of the identification and quantification results of the proteome and phosphoproteome in EGFR-TKI-sensitive (PC9) and EGFR-TKI-resistant (CL68) lung cancer cells in three biological triplicates. b Pearson’s correlation of phosphosite abundance in biological triplicate analysis. KEGG signaling pathways enriched (p < 0.05) from c upregulated and d downregulated phosphosites/phosphoproteins using STRING (two-sampled t-test, FDR < 0.01, S0 = 0.1). e Overall, 161 phosphosites and 49 proteins were quantified in the EGFR-TKI resistance signaling pathway, revealing differentially expressed proteins (two-sampled t-test, FDR < 0.01, S0 = 0.4) and phosphosites (two-sampled t-test, FDR < 0.01, S0 = 0.1). f Kinase motfi enrichment analysis using Fisher’s exact test (FDR < 0.02) where top ones shown among 30 kinase motifs. g Motif logo for ERK1, 2, and PKA or PKC kinases extracted by pLOGO (https://plogo.uconn.edu/). h Tyrosine phosphosites showing differential expression levels between the two cell lines (two-sampled t-test, FDR < 0.01, S0 = 0.1). Source data are provided as a Source Data file.