Fig. 2.
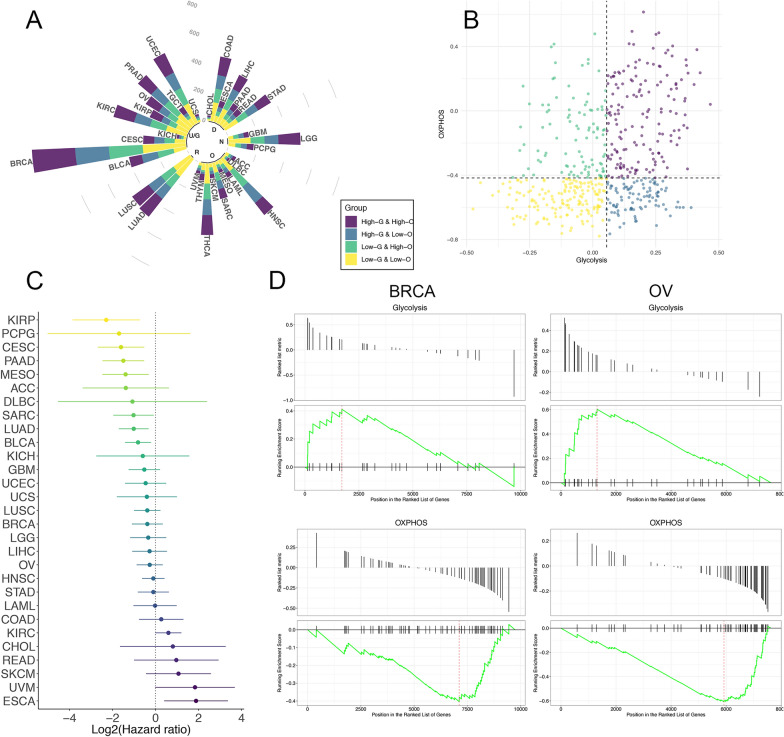
A The circular bar plot displays the percentages and numbers of samples in four different metabolic subgroups across multiple cancer types. The cancers are also annotated with the physiology systems they belong to: ‘R’, Respiratory system; ‘U/G’, Urinary system and Genital system; ‘D’, Digestive system; ‘N’, Nervous system; ‘O’, Other systems. B The scatter plot showing the distribution of Glycolysis score (x-axis) and OXPHOS score (y-axis) in tumor patients. Here we took lung adenocarcinoma as an example to illustrate the rule for subgroup classification: patients were assigned to four metabolic subgroups according to the median value of the two scores. C The prognostic value of metabolic classification (LGHO vs HGLO) in each cancer type, shown in the forest plot with corresponding hazard ratio (HR, log-transformed) and 95% confident interval (95%CI, log-transformed). D Gene set enrichment analysis (GSEA) based on mass proteomic spectrum data for BRCA and OV indicated that the proteins in the Glycolysis signature were significantly enriched in HGLO group, while proteins in OXPHOS signature were enriched in LGHO group