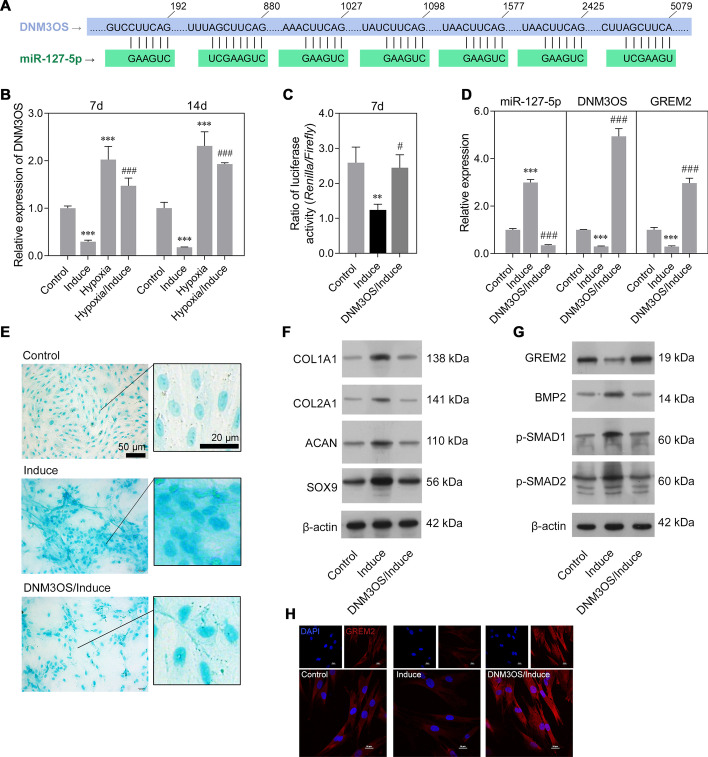
Fig. 6.
DNM3OS inhibited chondrogenic differentiation of rMSCs via miR-127-5p regulation of GREM2. The rMSCs were transfected with a DNM3OS overexpression plasmid, followed by induction of chondrogenic differentiation for 14 days. Accordingly, the rMSCs were classified into control, induction, hypoxia, hypoxia/induction, and DNM3OS/induction groups. A The putative binding site for miR-127-5p in the DNM3OS 3′-UTR is shown. B DNM3OS expression was determined by quantitative real-time PCR analysis. C A luciferase reporter assay was performed to identify direct action between DNM3OS and the GREM2 3ʹ-UTR. D Levels of DNM3OS, miR-127-5p, and GREM2 expression were determined by quantitative real-time PCR analysis. Results are expressed as mean ± standard deviation of data obtained from three independent experiments. **p < 0.01, ***p < 0.001, compared with control; #p < 0.05, ###p < 0.001, compared with induction group; E Images from Alcian blue staining assays of rMSCs. (F) Levels of COL1A1, COL2A1, SOX9, and ACAN protein expression were detected in rMSCs. G Levels of GREM2, BMP2, p-SMAD1, and p-SMAD2 in rMSCs were detected by western blotting. H GREM2 protein expression in rMSCs was measured by immunofluorescence