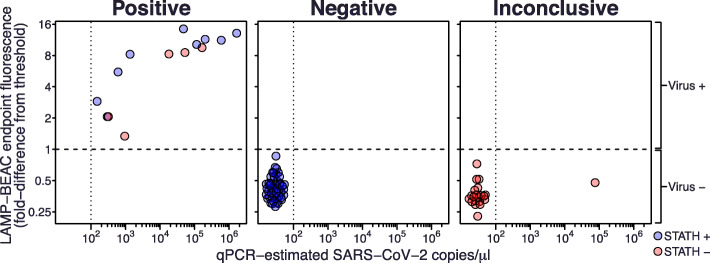
Fig. 4.
Validation of multiplexed LAMP-BEAC on 82 clinical saliva samples. Inactivated saliva samples were assayed by LAMP-BEAC using molecular beacons STATH_LBMB_S2, N2_LBMB_S3, and Penn_LFMB_S1. Samples were called “Positive” if they had detectable amplification in any SARS-CoV-2 amplicon; if human control STATH amplification was detected but not SARS-CoV-2, they were called “Negative”; if no STATH or SARS-CoV-2 amplification was detected they were called “Inconclusive.” SARS-CoV-2 targeted N2 and Penn fluorescence were quantified as the fold difference from a threshold set at two times the highest fluorescence observed in negative controls (dashed line). The maximum Penn or N2 fluorescence observed among the 5 reactions for each sample (y-axis) was compared to viral copy numbers estimates by laboratory qPCR (x-axis). Samples with qPCR copy numbers below the inferred limit of detection of 100 copies per μl are arbitrarily spaced apart for visualization. Sample integrity was assessed as STATH end point fluorescence greater than 120% the greatest fluorescence observed in water controls. Blue points indicate samples with detected STATH amplification and red indicates samples with no detectable STATH amplification, i.e., potentially indicating degraded samples or competition between amplicons