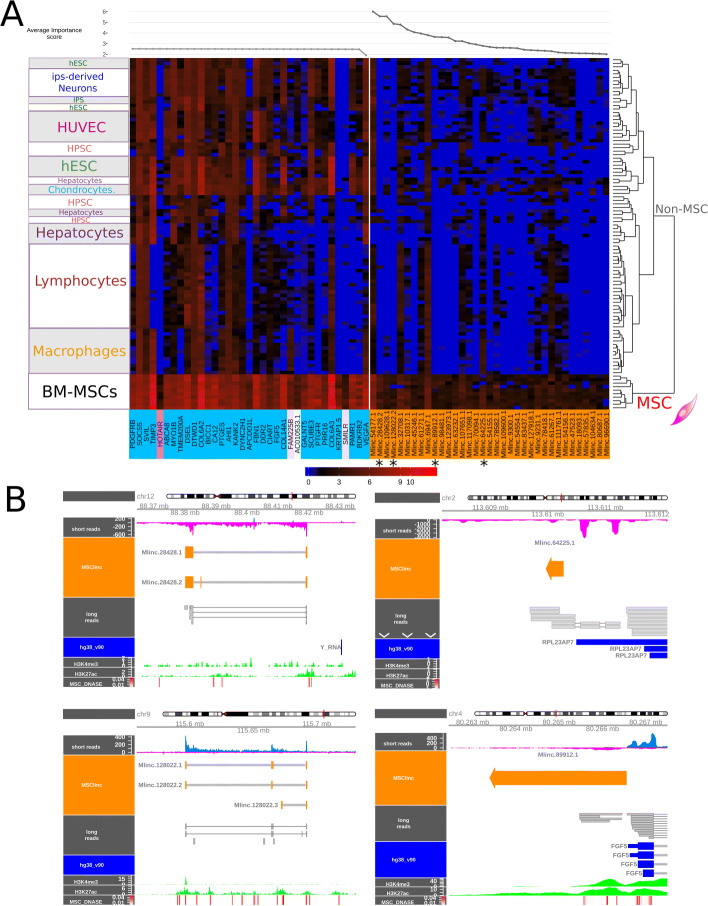
Fig. 3.
Selection of a refined set of the best candidates by random forest (top35), long-read sequencing and epigenetic features. a Expression of the best MSC-specific candidates selected by Boruta machine learning along MSC group and not MSC cohorts. Left pannel: top35 most relevant annotated genes (non-coding included); Right pannel: unannotated intergenic lncRNAs (Mlincs) and their average importance scores determined by Boruta method displayed in upside line plot. b Genomic visualisation of Mlincs 28428 (up left panel), 64225 (up right panel), 128022 (down left panel), and 89912 (down right panel). Predictions (Mlinc orange) from short reads alignment of all MSC group files (blue/magenta and BAM visualisation), are compared with unoriented long-read alignments (grey). Additional epigenomic features are shown to reveal active transcriptional activity from trimethylation of Histone H3 (H3K4me3), acetylation of Histone H3 H3K27 in MSCs (H3K4me3 and H3K27ac, green), and Dnase sensibitity hotspots of MSC (MSC DNAse, red)