Fig. 5.
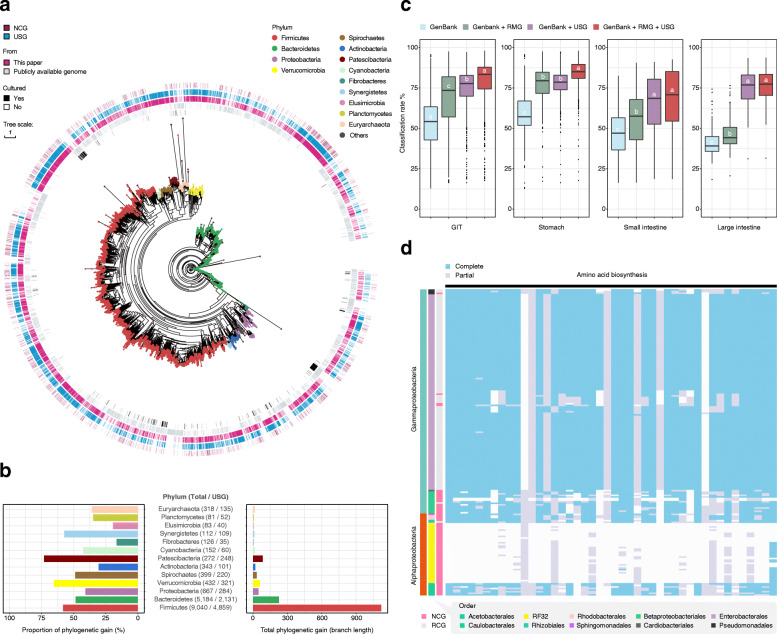
Novel species have distinct functional capacities. a The maximum-likelihood tree of the 10,373 MAGs identified in this study and previously published ruminant microbial genomes (17,425 genomes in total) was produced from concatenated protein sequences using PhyloPhlAn [30]. Clades are colored according to phyla. Genome information is presented in the outer layers. NCG, near-complete USGs. b The phylogenetic gain contributed by the microbial tree of the ruminant GIT provided by the USGs is shown as proportional increases in branch length per phylum (left) and absolute branch lengths (right). Numbers of USGs are indicated in parentheses (left, total; right, USGs). c Comparison of the read classification rates of all the ruminant GIT metagenomic samples using the following datasets: a common database consisting of all complete microbial genomes in GenBank, the GenBank database plus the RMG, the GenBank database plus the USGs identified in this study, and all three datasets. The Wilcoxon rank-sum test was used to assess the differences, and significant differences (P < 0.05) are indicated by different letters (a, b, c, and d). d Comparison of functional differences in genome properties between the NCG (n = 69) and RCG (n = 125) sets of Proteobacteria. The “Complete” and “Partial” heatmaps indicate functional profiles of GPs of the genomes involved in amino acid biosynthesis