Figure 2. Generation of GATA6−/− pluripotent stem cell lines.
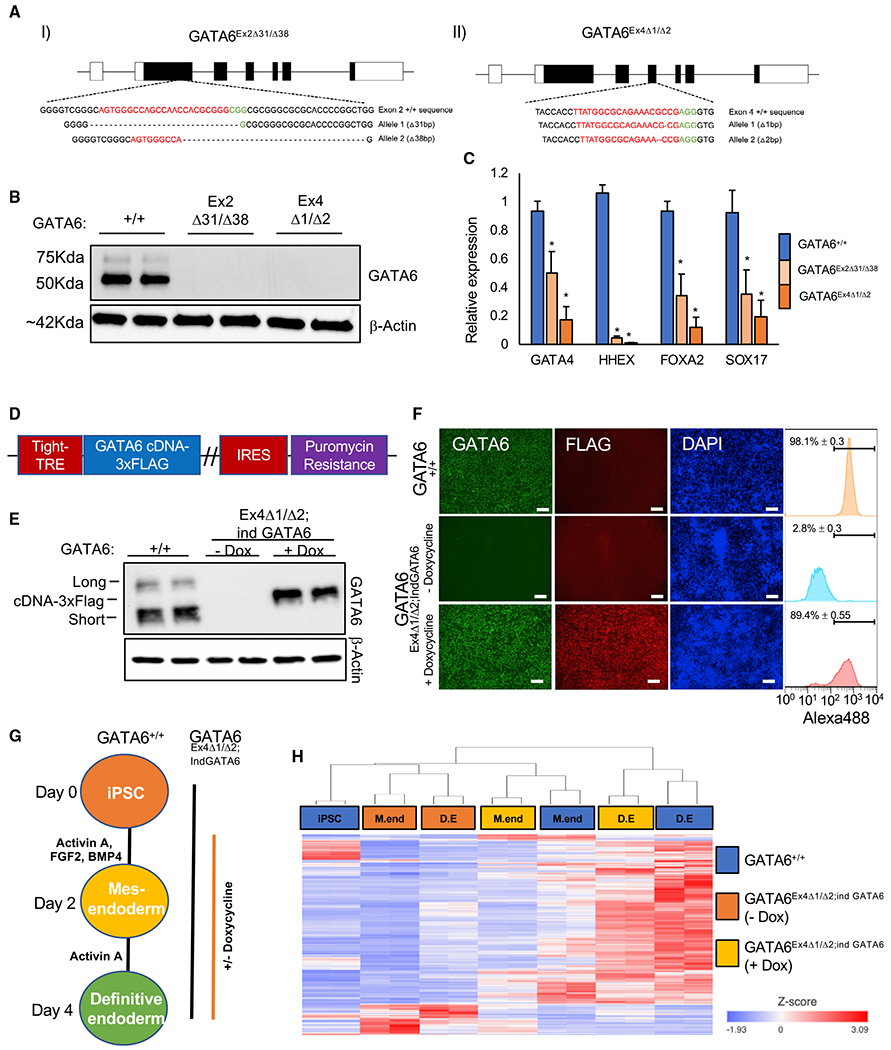
(A) CRISPR-Cas9-derived indels in K3 pluripotent stem cells using guide RNAs (gRNAs) targeting (I) exon 2 and (II) exon 4 of the GATA6 gene. Red denotes gRNA sequence; green denotes PAM sequence.
(B) Western blot analysis of GATA6 and β-actin in GATA6+/+, GATA6Ex2Δ31/Δ38 and GATA6Ex4Δ1/Δ2 cell lines at day 2 of differentiation.
(C) Graph showing qRT-PCR analysis of definitive endoderm-enriched mRNAs in definitive endoderm normalized to the housekeeping mRNA RPL13a. Mean values, n = 3, mean ± SD. Student’s t test: *p < 0.05.
(D) Doxycycline-inducible GATA6-3xFLAG cDNA plasmid.
(E) Western blot analysis of GATA6 and β-actin protein expression in GATA6+/+ and GATA6Ex4Δ1/Δ2;ind GATA6 pluripotent cells with and without doxycycline at day 2 of differentiation. Long, short, and exogenous short 3× FLAG epitope-tagged GATA6 isoforms are indicated.
(F) Immunofluorescence analysis of GATA6, FLAG, and DAPI in GATA6+/+ and GATA6Ex4Δ1/Δ2;ind GATA6 cells with and without doxycycline at day 2 of differentiation. Scale bars, 100 μm. GATA6-positive population: flow cytometry analysis, n = 3, mean ± SD.
(G) Differentiation protocol used to analyze GATA6 function.
(H) Heatmap with hierarchical clustering of GATA6-dependent genes during definitive endoderm formation in GATA6+/+ and GATA6Ex4Δ1/Δ2;ind GATA6 pluripotent cells with and without doxycycline. GATA6-dependent genes: false discovery rate (FDR) < 0.05: GATA6+/+ versus GATA6Ex4Δ1/Δ2;ind GATA6 (—dox), and GATA6Ex4Δ1/Δ2;ind GATA6 (—dox) versus GATA6Ex4Δ1/Δ2;ind GATA6 (+dox) at days 2 and 4 using gene-specific analysis (GSA; RNA-seq, n = 2).
