Abstract
Horizontal gene transfer (HGT) is a potentially critical source of material for ecological adaptation and the evolution of novel genetic traits. However, reports on posttransfer duplication in organism genomes are lacking, and the evolutionary advantages conferred on the recipient are generally poorly understood. Sucrase plays an important role in insect physiological growth and development. Here, we performed a comprehensive analysis of the evolution of insect β-fructofuranosidase transferred from bacteria via HGT. We found that posttransfer duplications of β-fructofuranosidase were widespread in Lepidoptera and sporadic occurrences of β-fructofuranosidase were found in Coleoptera and Hymenoptera. β-fructofuranosidase genes often undergo modifications, such as gene duplication, differential gene loss, and changes in mutation rates. Lepidopteran β-fructofuranosidase gene (SUC) clusters showed marked divergence in gene expression patterns and enzymatic properties in Bombyx mori (moth) and Papilio xuthus (butterfly). We generated SUC1 mutations in B. mori using CRISPR/Cas9 to thoroughly examine the physiological function of SUC. BmSUC1 mutant larvae were viable but displayed delayed growth and reduced sucrase activities that included susceptibility to the sugar mimic alkaloid found in high concentrations in mulberry. BmSUC1 served as a critical sucrase and supported metabolic homeostasis in the larval midgut and silk gland, suggesting that gene transfer of β-fructofuranosidase enhanced the digestive and metabolic adaptation of lepidopteran insects. These findings highlight not only the universal function of β-fructofuranosidase with a link to the maintenance of carbohydrate metabolism but also an underexplored function in the silk gland. This study expands our knowledge of posttransfer duplication and subsequent functional diversification in the adaptive evolution and lineage-specific adaptation of organisms.
Keywords: horizontal gene transfer, gene duplication, β-fructofuranosidase, insect, adaptation
Introduction
Horizontal gene transfer (HGT), also called lateral gene transfer, is the movement of genetic material across distantly related species outside of reproduction (Keeling and Palmer 2008; Keeling 2009). The greater prevalence and our deeper understanding of gene transfer in prokaryotes generally overshadow the importance of HGT in eukaryotes (Keeling and Palmer 2008). A mobile functional genetic element can be effectively introduced into a eukaryotic genome; however, it needs to overcome considerable evolutionary barriers, including entering the nucleus within the isolated germ cell line and adjusting to the transcription machinery of the recipient (Wybouw et al. 2016). Recently, there has been a growing incidence of HGT cases involving the movement of genetic information between prokaryotic symbionts and their eukaryotic hosts and even between eukaryotes (Dunning Hotopp et al. 2007; Soucy et al. 2015). However, only a few HGT events have been reported to be advantageous, and most genes transferred in HGT events have neutral or slightly detrimental effects; the genes gradually degenerate or lose their original functions in the recipient (Gogarten and Townsend 2005; Husnik and McCutcheon 2018). Nevertheless, the number of well-supported cases of HGT, many with underlying functional or ecological implications for their new host, is now increasing rapidly (Danchin et al. 2010; Acuña et al. 2012; Husnik and McCutcheon 2018; Ren et al. 2020; Wang et al. 2020). For example, several transferred genes might effectively participate in the phenotypic divergence and biochemical metabolism of insect species, which implies that HGT may have the potential to influence the functional evolution of eukaryotic recipients (Li et al. 2011; Zhu et al. 2011; Sun et al. 2013; Danchin et al. 2016). However, direct physiological evidence shedding light on the detailed physiological mechanisms supporting this issue has been scarce.
Gene duplication is considered integral to the origin of novel traits because it offers raw genetic material for evolution and the subsequent generation of diverse biological functions (Helmkampf et al. 2015; Turetzek et al. 2016; Kursel and Malik 2017; Castagnone-Sereno et al. 2019; Dyson and Goodisman 2020). Gene duplication occurs more commonly among horizontally transferred genes than among endogenetic genes (Li et al. 2011). However, newly duplicated genes tend to have redundant functions immediately, and one of the copies could accumulate deleterious mutations to generate pseudogenes (pseudogenization) and eventually be lost without any effect on the survival fitness of an individual (Assis and Bachtrog 2013). However, the dosage effect produces a selective advantage when the duplicated genes are functional redundantly or divergently, ensuring that both copies are retained in the genome and evolve under purifying selection after a possible period of relaxed constraint. Functional divergence of the duplicated genes can occur through neofunctionalization or subfunctionalization (Fan et al. 2008; Kursel and Malik 2017; Heidel-Fischer et al. 2019; Wang et al. 2019; Zhou et al. 2019). Therefore, investigating the subsequent fate of these duplicate gene pairs will contribute to the understanding of the key role of HGT in the evolution of eukaryotes.
Lepidoptera (butterflies and moths) is the second-largest order of insects, comprising nearly 160,000 described extant species (Li et al. 2018). Compared with other insect orders, diverse types and a considerable number of HGT events have been observed in lepidopteran insects (Wheeler et al. 2013). Previous studies found that 14 types of 22 genes were detected as candidate HTGs in several lepidopteran insects, and more than half appear to have undergone duplication events after their acquisitions (Li et al. 2011). Of these, β-fructofranosidase (β-FFase, EC 3.2.1.26), glycerophosphoryl diester phosphodiesterase, nicotinamide adenine dinucleotide-dependent epimerase/dehydratase, and cysteine synthase were duplicated, and multiple copies have been maintained in several lepidopteran genomes after HGT (Li et al. 2011; Zhu et al. 2011; Sun et al. 2013). However, the underlying functional innovation and adaptation to lepidopteran hosts in their ancient lineages remain largely unexplored.
Sucrose is a major carbohydrate reserve and is the most commonly transported sugar in host plants, providing a rich source of carbon and energy for insect metabolism (Li et al. 2017). Insect sucrase activity depends principally on α-glucosidases; however, the sucrose hydrolases of Lepidoptera occur as three distinct forms of enzymes: α-glucosidases, sucrose hydrolases (SUHs), and β-fructofranosidases (Lee et al. 2012; Wang et al. 2015; Li et al. 2017). β-fructofranosidase, which has been described thoroughly in bacteria, fungi, and plants, and identified in several lepidopteran insects as a result of HGT from bacteria (Hirayama et al. 2007; Daimon et al. 2008; Gan, Li, et al. 2018; Guo, Wang, et al. 2018). As shown in recent studies, β-fructofuranosidase genes exist both in the lepidopteran insects and in the coleopteran insects (Keeling et al. 2013; Pedezzi et al. 2014; Zhao et al. 2014). We discovered that these insect species appear to experience specific β-fructofranosidase gene duplications after HGT from bacteria. We found that multiple copies of β-fructofranosidases were retained in their genomes. However, comprehensive evolutionary analysis of β-fructofranosidases in insects is lacking, and the diverse fates of β-fructofranosidase duplicates formed by posttransfer duplication remain largely unknown.
In the present study, we systematically performed genomic analyses, including sequence presence/conservation and chromosomal synteny, and reconstructed the evolutionary relationships between the members of β-fructofuranosidase gene family in insects. Our analysis indicated that gene duplication events gave rise to multiple lepidopteran-specific β-fructofuranosidase (named SUC in Lepidoptera), and gene loss subsequently occurred in several lepidopteran species. We integrated data regarding expression and enzymatic properties analyses to examine the functional divergence of SUCs in Lepidoptera. Moreover, our study focused on functional analysis of the single functional ortholog in Bombyx mori, called BmSUC1, with the goal of better understanding its physiological functions in the absence of genetic redundancy. We reported insights from both phenotypic observations and transcriptome investigations into the underlying physiological functions of SUC. Based on our results, the universal function of SUC in the midgut was uncovered, and an underexplored space of SUC gene function in the silk gland was examined. The present study provides significant insights into the evolutionary contributions of HGT and the subsequent functional diversification to organismal adaptation in eukaryotes.
Results
Gene Duplication of β-Fructofuranosidases in Insects
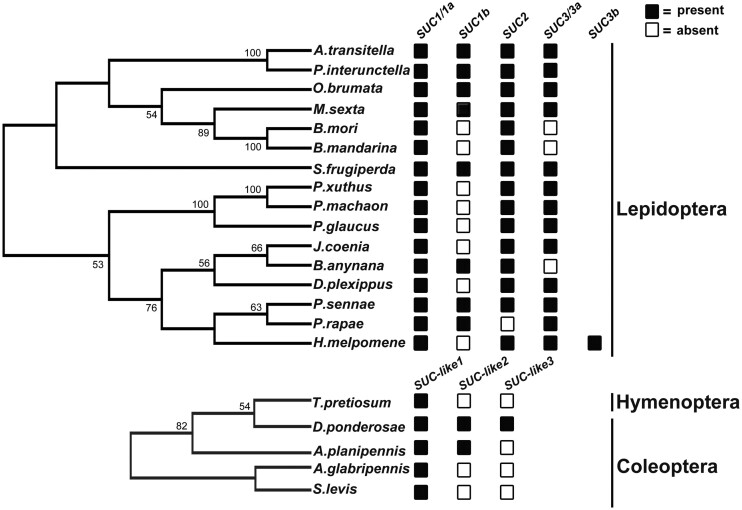
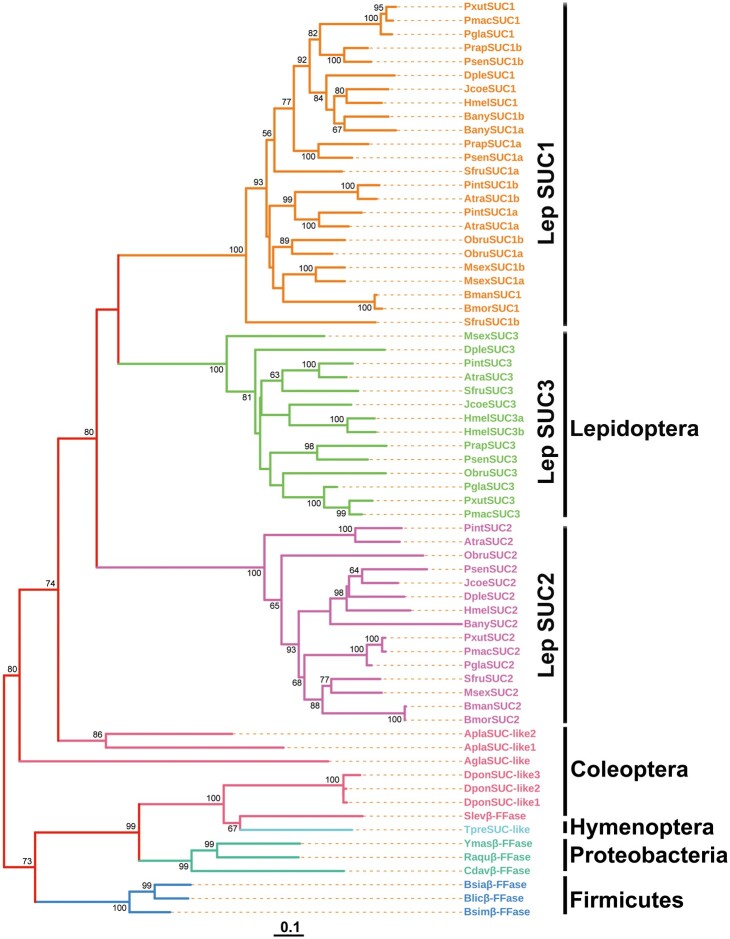
To understand the evolutionary history and phylogenetic relationships of β-fructofuranosidase encoding genes in insects, we systematically examined the distribution of SUC and its homologs among the insect high-quality genomes using B. mori SUC1 and SUC2 as queries (supplementary table S1, Supplementary Material online). We identified SUC and SUC homologous proteins in 21 insect genomes: 16 lepidopteran, four coleopteran, and one hymenopteran species. We performed a TBlastN search against 54 hymenopteran genomes in the RefSeq collection of the National Center for Biotechnology Information (NCBI). Consequently, we only found one homolog in Trichogramma pretiosum among the 54 hymenopteran species that we surveyed. β-fructofuranosidase was encoded by a single gene in T. pretiosum. In coleopteran species, Anoplophora glabripennis and Sphenophorus levis contain one SUC homologous protein, whereas Agrilus planipennis and Dendroctonus ponderosae have two and three SUC homologous proteins, respectively. Intriguingly, we found three or more SUC paralogs in several lepidopteran insects (fig. 1). Among these, the largest number of SUC homologous copies (SUC1a, SUC1b, SUC2, and SUC3) was observed in five moth species: Operophtera brumata, Manduca sexta, Spodoptera frugiperda, Plodia interpunctella, and Amyelois transitella. Two or three SUC copies were observed in the genomes of B. mori, Bombyx mandarina, Pieris rapae, Papilio glaucus, Papilio machaon, Papilio xuthus, Junonia coenia, Bicyclus anynana, and Danaus plexippus (fig. 1). We then conducted a phylogenetic study via the maximum-likelihood (ML) method based on the examined species to further identify the evolutionary relationships of SUC and SUC homologous proteins. We found that lepidopteran SUCs were clustered into three distinct, highly supported clades, which suggested that these proteins had diverse functions before the divergence of lepidopteran species. SUC1a was grouped with SUC1b and formed a sister clade with SUC3, suggesting that SUC1a/1b was likely formed by gene duplication in these lepidopteran genomes. Moreover, we found that SUC2s from the 15 lepidopteran species formed a monophyletic clade and are likely orthologous (fig. 2).
Fig. 1.
Distribution of SUC paralogs across lepidopteran species. Summary of SUC homolog presence across Lepidoptera with two expanded orders: Coleoptera and Hymenoptera. A total of 61 full-length genes encoding putative β-fructofuranosidases from 21 insect species were detected. The presence (black box) or absence (white box) of each SUC and SUC-like were identified by retrieving the available coding sequences from NCBI, Ensembl, and Lepbase online databases is schematized to the right of each species. Three conserved mitochondrial genes (cytochrome oxidase subunit I, cytochrome oxidase subunit II, and NADH dehydrogenase subunit 1) were used to build the evolutionary tree of lepidopteran species. Cytochrome oxidase subunit I genes were used to build the phylogenetic relationship between hymenopteran and coleopteran species. Phylogenetic trees were constructed with multilocus sequence analysis (MLSA) using the IQ-TREE software. The highest log likelihood is −12801.62 and −2570.39, respectively. Detailed information involved sequences and the accession numbers are provided as supplementary tables S1 and S4, Supplementary Material online.
Fig. 2.
Evolutionary relationship among SUCs. The evolutionary relationship was constructed using MEGA7.0 software with the maximum-likelihood topology based on the (LG+G+I) model. The analysis involved 67 amino acid sequences. All positions containing gaps and missing data were eliminated. Bootstrap values greater than 60 are shown. The scale bar represents 0.1 amino acid substitutions per site. Sequences and the accession numbers are provided in supplementary table S1, Supplementary Material online.
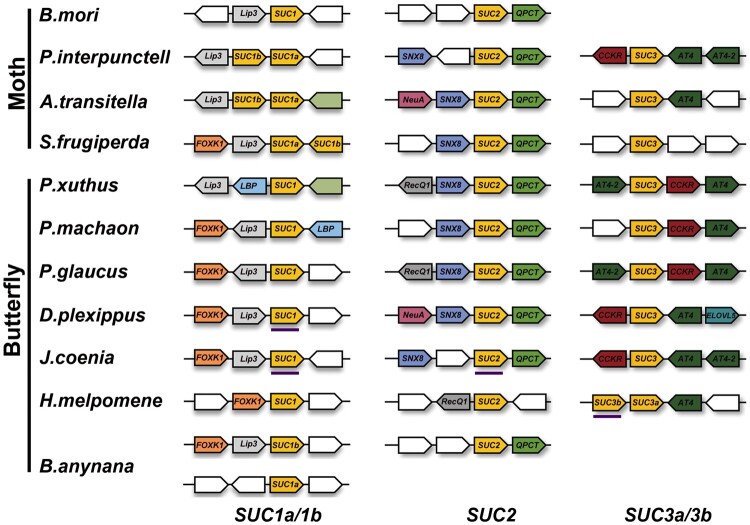
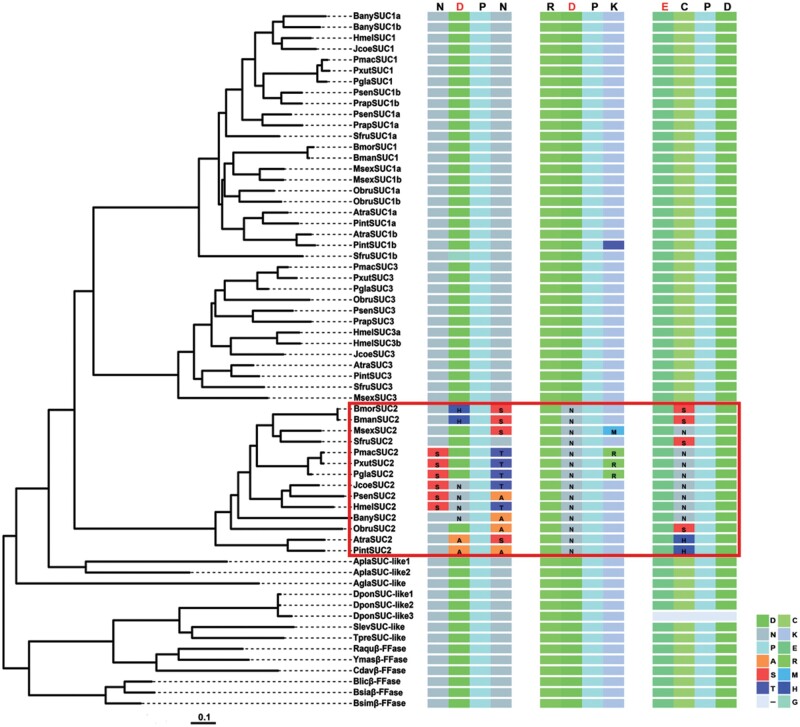
The synteny, order, and orientation surrounding the SUC locus were relatively well conserved among the 11 lepidopteran insect genomes surveyed. However, S. frugiperda SUC3 and B. anynana SUC1a represented two interesting exceptions (fig. 3). Of note, we found SUC1a located in close proximity to SUC1b, and it was tandemly arranged on the same scaffold in several species, including S. frugiperda, P. interpunctella, and A. transitella (fig. 3). The unique syntenic location, together with the above phylogenetic tree, further suggested that two gene copies, SUC1a and SUC1b, were formed via a gene duplication event in these lepidopteran species. The gene duplication event appears to be true for SUC3 in Heliconius melpomene, where two adjacent copies (SUC3a and SUC3b) were found in the genome. Among the insects examined, we discovered that a great majority of SUC homologs were intron-less, implying that the gene structures of insect β-fructofuranosidases were highly conserved during evolution (fig. 3). We then performed further multiple alignments to annotate evolutionarily or structurally related sites or regions among lepidopteran SUC sequences. We found that the signatures of catalytically active β-fructofuranosidase, that is, Asp63 (D), Asp181 (D), and Glu234 (E), were conserved in SUC1s and SUC3s. Furthermore, they shared a high sequence similarity and conserved substrate binding sites (supplementary fig. S1, Supplementary Material online). By contrast, three catalytic regions (62–65,180–183, and 234–237 amino acids; numbered according to BmSUC1) of β-fructofuranosidase are not conserved in SUC2s, which are unlikely to be functional β-fructofuranosidases (fig.4 and supplementary fig. S1, Supplementary Material online).
Fig. 3.
Conserved syntenic analysis of SUCs and their surrounding genes. Synteny was analyzed using PxSUC1, PxSUC2, and PxSUC3 as anchor sites, respectively. Lepidopteran SUCs are placed in the middle and indicated in yellow. Homologs are represented by the same color, and genes with no homolog are indicated with blank boxes. Forkhead box protein k1, FOXK1; allatostatin-A receptor, AstAR; lipopolysaccharide-binding protein, LBP; glutaminyl-peptide cyclotransferase-like, QPCT; sorting nexin-8-like, SNX8; acylneuraminate cytidylyltransferase, NeuA; ATP-dependent DNA helicase Q1-like, RecQ1; amino acid transporter 4-like, AT4/AT4-2; cholecystokinin receptor-like, CCKR; elongation of very long-chain fatty acids protein 5: ELOVL5. The direction of the gene is indicated with block arrows. Gene name abbreviations are shown inside the arrow diagrams. The positions of genes on the chromosomes are not drawn to scale. SUC homologs with introns are underlined in purple: DpleSUC1, JcoeSUC1, JcoeSUC2, and HmelSUC3b.
Fig. 4.
Partial amino acid sequence alignment of SUCs. Three catalytic regions (62–65,180–183, and 234–237 amino acids; numbered according to BmSUC1) were extracted following an alignment of the full-length SUC protein sequences in MEGA7.0 program. The R package “ggtree” was used to visualize and annotate the result. Same amino acids are represented by the same color. Three conserved catalytic residues Asp63 (D), Asp181 (D), and Glu234 (E) are indicated in red letters. The alignment result of SUC2s is marked with a red rectangle. The scale bar represents 0.1 amino acid substitutions per site.
Expression Divergence of Duplicated Lepidopteran SUCs
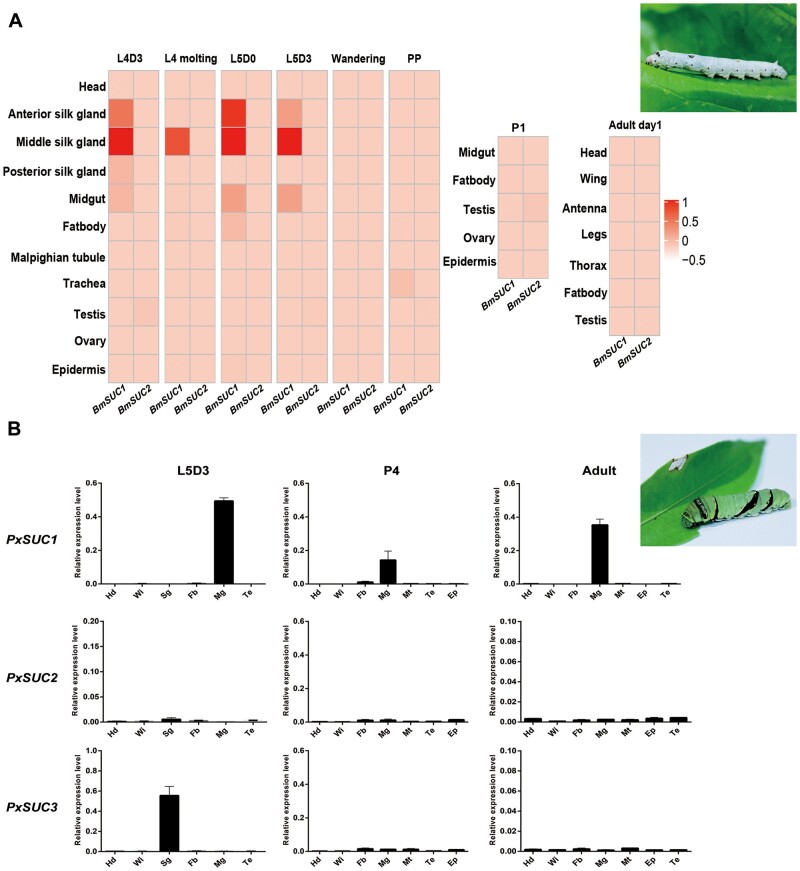
Given the coretention of SUC paralogs in lepidopteran genomes, we considered the possibility that these paralogs developed specialized function, potentially through tissue-specific expression patterns. Bombyx mori is a model lepidopteran insect, and its high-quality reference genome is conducive to our further research (Xia et al. 2004). We then determined any evidence for tissue-specific expression by analyzing the previously published genome-wide RNA-seq data. We retrieved the data from SilkDB 3.0 and determined the temporal expression profiles of SUC paralogs across tissues and developmental stages in B. mori. The RNA-seq profiling revealed that the BmSUC1 transcript was primarily detected in the midgut but not in the head, fat body, Malpighian tubule, and testis (fig. 5A). Of note, the highest level of BmSUC1 expression was observed in both the anterior silk gland (Asg) and middle silk gland (Msg) but not the posterior silk gland (Psg). In the midgut and silk gland, the expression of BmSUC1 was mainly detected during the feeding stage (L4D3, L5D0, and L5D3), but disappeared by the wandering stage, just prior to the start of spinning, pupal, and adult stages (fig. 5A). Consistent with previous studies, the expression of BmSUC2 was very low in any of the tissues or developmental stages examined (fig. 5A and supplementary fig. S2A, Supplementary Material online) (Daimon et al. 2008; Guo, Wang, et al. 2018). We then extended our expression analysis of SUC paralogs to other lepidopteran species that contain duplicate SUC genes. The spatial and temporal expression profiles of SUC were examined in the butterfly, P. xuthus (Lepidoptera: Papilionidae), which was also a good model lepidopteran insect. We found that PxSUC1 was highly expressed in the larval midgut and dropped markedly in the pupal midgut. Of note, the PxSUC1 expression level was remarkably increased in the adult midgut (fig. 5B). Western blotting confirmed that strong signals for PxSUC1 were universally observed throughout the entire midgut (supplementary fig. S2B and C, Supplementary Material online). Furthermore, the PxSUC1 protein was predominantly localized in the goblet cells of the midgut (supplementary fig. S2D, Supplementary Material online). By contrast, the relatively high level of PxSUC3 expression was primarily restricted to the larval silk gland compared with the other developmental stages or tissues, including the head, midgut, fat body, and Malpighian tubules (fig. 5B). PxSUC2 expression was extremely low, which was similar to that of BmSUC2 (fig. 5B). Taken together, SUC copies display significantly divergent expression patterns in two species, indicating that the underlying functional differentiation probably occurred among lepidopteran SUCs.
Fig. 5.
Expression patterns of SUCs in Bombyx mori and Papilio xuthus. (A) Expression patterns of BmSUCs in B. mori. Expression data from SilkDB 3.0 (https://silkdb.bioinfotoolkits.net/main/species-info/-1, last accessed March 21, 2021) were searched to identify the expression profiles of SUC genes in 78 samples throughout the silkworm development stages (larval, pupal, and adult stages). The fold-changes of SUCs were indicated by the R package “Complex Heatmap.” (B) qRT–PCR analysis of PxSUCs expression. Samples were collected from the dissections of the head (Hd), wing disc (Wi), silk gland (Sg), fat body (Fb), midgut (Mg), Malpighia tubule (Mt), testis (Te), and epidermis (Ep). qRT–PCR was normalized using Pxrpl as a control. Values represent means ± SEM (n = 3). L4D3, the third day of the fourth larval instar; L4 molting, the end of the fourth larval instar; L5D0, the start of the fifth larval instar; L5D3, the third day of the fifth larval instar; PP, prepupal stage; P1, the first day of the pupal stage; P4, the fourth day of the pupal stage; Adult, adult stage; Adult day1, the first day of the adult stage.
Biochemical Characteristics Divergence of Three SUC Proteins
Although we detected the expression differentiation of SUC genes among lepidopteran species, it is unclear whether the functions of these genes were divergent during evolution. Three recombinant PxSUC proteins were purified and the substrate specificities were examined by testing several sugar substrates (sucrose, maltose, and isomaltose) to evaluate the evolutionary changes in duplicate gene pairs (supplementary figs. S3 and S4, Supplementary Material online). PxSUC1 displayed high enzymatic activity toward sucrose but not maltose or isomaltose. PxSUC3 enzyme exhibited similar substrate specificities to PxSUC1, whereas PxSUC3 showed a lower hydrolytic activity toward sucrose than PxSUC1 (fig. 6A). These results suggested that PxSUC3 was not a pseudogene and was retained to perform a sucrose hydrolase function. In addition, it was interesting to notice that PxSUC2 had no hydrolytic activity toward any of the three sugar substrates, supporting the hypothesis that PxSUC2 turned into a nonfunctional β-fructofuranosidase (fig. 6A). Because purified PxSUC2 protein lacked enzymatic activity, only PxSUC1 and PxSUC3 proteins were analyzed further. The pH curve revealed that the optimum pH for PxSUC1 was approximately 7.0, whereas the enzymatic activity of PxSUC1 decreased by approximately 60% at pH 5.0 (fig. 6B). By contrast, the optimal pH for PxSUC3 was approximately 6.0, and it retained 80% of its maximum enzymatic activity at pH 5.0, suggesting that PxSUC3 is possibly an acid hydrolase (fig. 6B). We determined the kinetic constants by using sucrose as a reaction substrate to further explore the enzymatic activities of the two SUCs. The calculated Vmax and Km values of PxSUC1 and PxSUC3 were 0.40 ± 0.02 and 0.18 ± 0.05 μmol/min/μg and 34.9 ± 0.2 and 98.3 ± 0.16 mM, respectively (supplementary fig. S5, Supplementary Material online). Taken together, these data indicate that the three PxSUC proteins have a clear divergence in enzymatic properties.
Fig. 6.
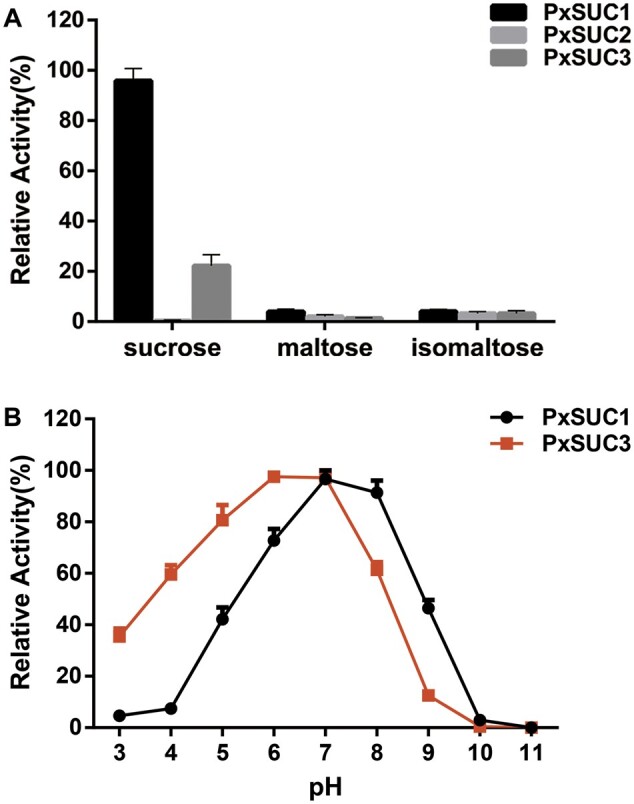
Enzymatic properties of three recombinant PxSUC proteins. (A) Enzymatic activities of recombinant PxSUC proteins. The purified PxSUC proteins were incubated with selected substrates (sucrose, maltose, and isomaltose). (B) The pH profiles of recombinant PxSUC1 and PxSUC3. The optimal pH was determined using sucrose as a substrate in 20 mM Britton–Robinson wide range buffer (pH: 3.0–11.0). Data represent the mean of three independent experiments, and error bars represent SDs.
BmSUC1 Plays an Important Role in the Development of Silkworm Larvae
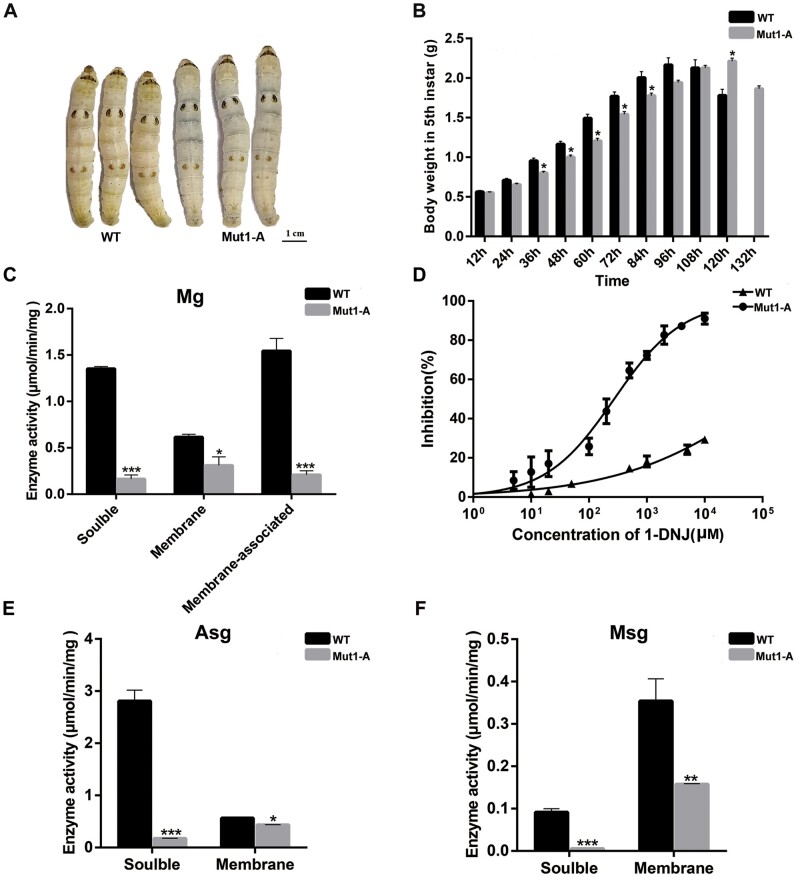
To ascertain the comprehensive physiological functions of SUC, we generated mutations that inactivated SUC1 in B. mori using CRISPR/Cas9 (supplementary fig. S6A, Supplementary Material online). Previous studies and the above results have shown that the active sites were absent in BmSUC2; hence, we genetically ablated BmSUC1 (Daimon et al. 2008). Two sgRNAs against the BmSUC1 gene were injected into the embryos (supplementary fig. S6A, Supplementary Material online). Of these, only sgRNA site1 was highly efficient. After seven generations of screening, two different types of chromosomal deletions around the target site one of BmSUC1 were detected: Mut1-A, Mut1-B, and Mut2 (supplementary fig. S6B, Supplementary Material online). Furthermore, a complete disappearance in BmSUC1 translational levels was observed in the midgut and silk gland tissues dissected from three mutant lines, demonstrating that this CRISPR/Cas9-induced mutagenesis system was effective (supplementary fig. S6C and D, Supplementary Material online). Compared with wild-type (WT) individuals, the mutants of BmSUC1 exhibited nonlethal phenotype and generated heritable targeted mutations, indicating that the interruption of BmSUC1 did not interfere with the viability or fertility of silkworms (supplementary fig. S7A and B, Supplementary Material online). Because three homozygous individuals showed similar phenotypes, we chose Mut1-A for subsequent experimental observations. First, we recorded the larval body weight in the fifth instar and observed discrepancies in the development rate from the second day of the fifth larval instar. In addition, the weights of mutant individuals were lower than that of controls (fig. 7A and B). Second, we found that the feeding intake of BmSUC1 mutants was generally less than that of WT (supplementary fig. S7C, Supplementary Material online). Moreover, compared with controls, BmSUC1 mutant larvae extended the duration of the final larval instar larval stage by approximately 12 h (fig. 7A and B). These findings unequivocally indicate that the interruption of BmSUC1 has adverse growth effects on B. mori larvae.
Fig. 7.
CRISPR/Cas9-mediated gene editing of BmSUC1 in Bombyx mori. (A) Individual body sizes on day 5 of the fifth larval instar. (B) The weight of each individual during the fifth instar (n = 20). (C) Sucrase activities of the midgut in WT and BmSUC1 mutant larvae. (D) Inhibitory effects of 1-deoxynojirimycin on midgut sucrase activities in WT and mutant. (E) Sucrase activities of anterior silk gland in WT and mutant larvae. (F) Sucrase activities of middle silk gland in WT and mutant larvae. Statistical analyses were performed using an independent Student’s t-test. *P < 0.05, **P < 0.01, and ***P < 0.001. Vertical bars indicate means ± SEM (n = 3).
BmSUC1 Is Required for Sucrase Activity and the Sugar-Mimic Alkaloid Resistance
We next performed biochemical analyses of midgut sucrases, including soluble, membrane-associated, and membrane-bound in WT and BmSUC1 mutants. As expected, the midgut sucrase activities of the BmSUC1 mutant line were markedly lower than those of WT (fig. 7C). Then, we extended the analyses to the Asg and Msg. The sucrase activities of WT were similarly higher than those of BmSUC1 mutants (fig. 7E and F). These results suggest that BmSUC1 participates in the digestion of sucrose and serves as a critical sucrase. 1-Deoxynojirimycin (1-DNJ) is a common component of mulberry latex and occurs in higher concentrations in mulberry than in other plants (Konno et al. 2006). 1-DNJ, as a potent α-glucosidase inhibitor, can competitively bind to α-glucosidase in the intestine and possesses a higher affinity than disaccharides, such as sucrose (Hirayama et al. 2007). We then examined the inhibitory activity of 1-DNJ on the midgut sucrase activities in the fifth instar larvae of the WT and BmSUC1 mutants. The midgut sucrase activity of mutants was strongly inhibited by 1-DNJ even at a low concentration. By contrast, the midgut sucrase activity of WT was much less affected by 1-DNJ (fig. 7D). Thus, these results indicate that the midgut sucrase activities of BmSUC1 mutants decrease and are more sensitive to 1-DNJ.
BmSUC1 Supports Larvae Metabolic Homeostasis
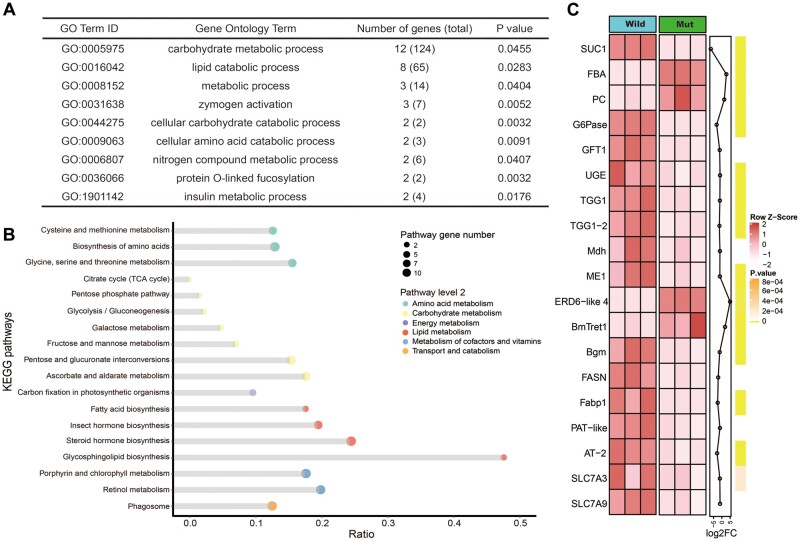
The decrease in the midgut sucrase activity and sensitivity to sugar-mimic alkaloid in BmSUC1 mutants raise the possibility that BmSUC1 was involved in sucrose metabolism in larvae. Thus, we performed the RNA-seq analysis of the midgut of controls and BmSUC1 mutants. RNA-seq analysis revealed broad effects of inactivated BmSUC1 on transcription, with 423 transcripts expressed at elevated levels and 462 transcripts expressed at reduced levels (supplementary fig. S8A, Supplementary Material online). Gene ontology (GO) enrichment analysis indicated that the significantly affected molecular functions were mainly involved in binding, transporter activity, and catalytic activity (supplementary fig. S8B and table S2, Supplementary Material online). In addition, GO analysis revealed significant enrichment of metabolic processes, with carbohydrate metabolism as a top category (fig. 8A). Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis showed that the significant enrichment pathways were mainly related to carbohydrate metabolism and energy metabolism (fig. 8B). Consistent with this, multiple genes encoding enzymes in fructose and mannose metabolism exhibited significant expression change in BmSUC1 mutants along with many genes in the glycolysis, pentose phosphate pathway, and tricarboxylic acid (TCA) cycle (fig. 8C). For instance, pyruvate carboxylase, which fuels the TCA cycle with one of its intermediates and participates in the first step of gluconeogenesis, was expressed at elevated levels in the midgut of BmSUC1 mutants. By contrast, glucose-6-phosphatase (G6Pase), which converts glucose-6-phosphate to glucose and is responsible for the homeostatic regulation of blood glucose levels, was significantly downregulated. Moreover, we found that most genes associated with pyruvate metabolism, galactose metabolism, and ascorbate and aldarate metabolism were expressed at reduced levels. Of note, trehalose transporter that mediated the bidirectional transfer of trehalose was upregulated (fig. 8C). These divergent effects on glycometabolism metabolisms may reflect the disruption of homeostatic glucose levels in the midgut of BmSUC1 mutants. In addition, multiple genes involved in the lipid metabolism, including long-chain-fatty-acid–CoA ligase bubblegum, fatty acid synthase, and fatty acid-binding protein 1 (Fabp1), were expressed at reduced levels in BmSUC1 mutants (fig. 8C). Furthermore, multiple amino acid transporters were also significantly downregulated in BmSUC1 mutants (fig. 8C). Taken together, these results indicate that the interruption of BmSUC1 affects internal metabolism that follows digestion and absorption, suggesting that BmSUC1 supports larval internal metabolic homeostasis.
Fig. 8.
Transcriptomic changes following interruption of BmSUC1 in the midgut. (A) Gene ontology (GO) enrichment analysis. GO terms with P < 0.05 were considered significant enrichment. (B) The KEGG pathways enrichment analysis. The different-sized dots represent the numbers of enriched genes in pathways. (C) Heat map of midgut DEGs. The color of heat map corresponds to the scale value of FPKM (row Z scores). The line chart displays the fold-change of DEGs. The green vertical bars represent the P value of each gene. The names of the DEGs are shown on the left. The detailed DEGs are provided in supplementary table S6, Supplementary Material online.
Functional HGT of SUC Contributes to Metabolic Homeostasis of Silk Gland
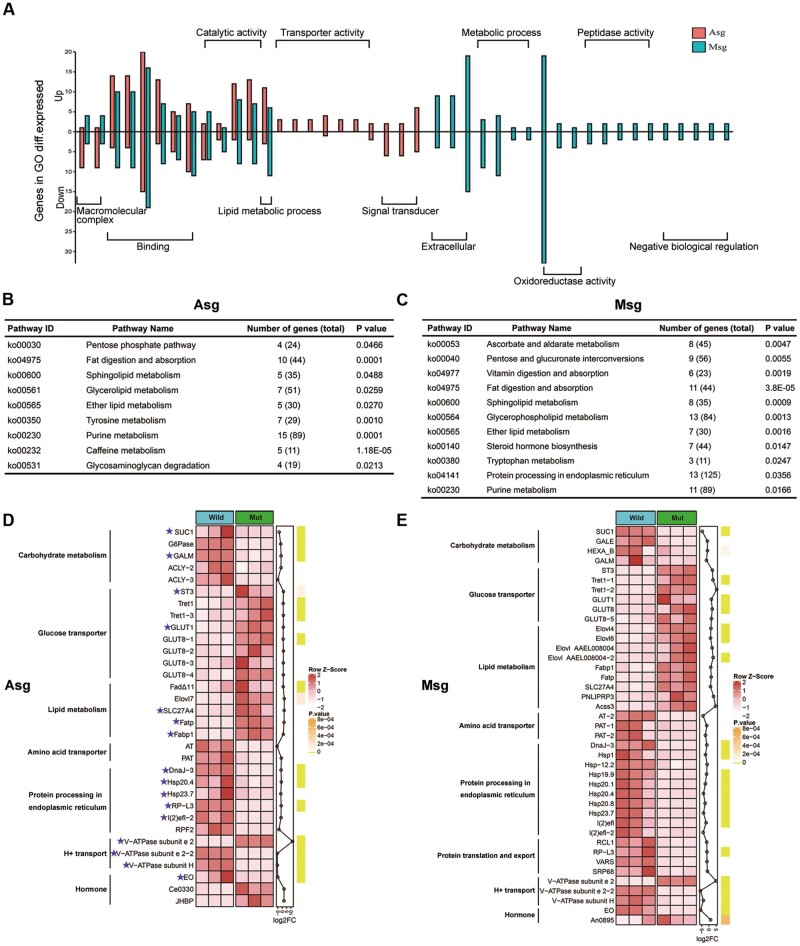
Expression profiles showed that SUC1 was mainly expressed in the anterior and middle silk gland tissue of B. mori. To obtain a molecular understanding of BmSUC1 function in silk gland, we performed the RNA-seq analysis using RNA isolated from Asg and Msg tissues on day 3 of the fifth instar in controls and BmSUC1 mutants. A total of 555 transcript levels were elevated and 307 transcript levels were reduced in Asg, whereas 424 and 447 transcripts were up- and downregulated in Msg, respectively (supplementary fig. S9A and B, Supplementary Material online). GO analysis of differentially expressed genes (DEGs) revealed significant enrichment of the macromolecular complex, binding, catalytic activity, and lipid metabolic process in both Asg and Msg (fig. 9A and supplementary table S3, Supplementary Material online). Intriguingly, GO terms, including metabolic process, oxidoreductase activity, extracellular, and peptidase activity, were only enriched in Msg. By contrast, GO terms for transporter activity and signal transducer were specifically enriched in Asg (fig. 9A). In addition, KEGG enrichment analysis of DEGs identified similar pathways activated in Asg and Msg, including carbohydrate metabolism, lipid metabolism, and amino acid metabolism (fig. 9B and C; supplementary fig. S10A and B, Supplementary Material online). Of note, pathways for protein processing in endoplasmic reticulum were specifically enriched in Msg (fig. 9C).
Fig. 9.
Transcriptomic changes following interruption of BmSUC1 in the silk gland. (A) The up- and downregulation of DEGs within enriched GO terms. Red/green bars represent the numbers of genes that were significantly up- and downregulated in Asg and Msg, respectively. The detailed GO terms information are described in supplementary table S3, Supplementary Material online. (B) The KEGG pathways enrichment analysis in Asg. (C) The KEGG pathways enrichment analysis in Msg. Pathways with P < 0.05 were considered as significant enrichment. (D) The heat map of selected DEGs in Asg. (E) The heat map of selected DEGs in Msg. The same DEGs in Asg and Msg are marked with purple asterisks. The color of heat map corresponds to the scale value of FPKM (row Z scores). The green vertical bars represent the P value of each gene. The detailed DEGs are provided in supplementary table S7, Supplementary Material online.
Among DEGs, 536 and 546 genes were specifically expressed in Asg and Msg, respectively, and shared 326 DEGs (supplementary fig. S9A, Supplementary Material online). These DEGs act in several metabolic pathways, including carbohydrate metabolism, amino acid metabolism, and fatty acid metabolism. Four DEGs related to carbohydrate metabolism were expressed at reduced levels in Asg, including G6Pase, aldose 1-epimerase (GALM), and two ATP citrate (pro-S)-lyases. Likewise, three DEGs for GALM, beta-hexosaminidase subunit beta, and UDP-glucose 4-epimerase, which were associated with the carbohydrate metabolism, also showed significant downregulation in Msg (fig. 9D and E). By contrast, multiple trehalose transporters and insulin-regulated facilitative hexose transporters that mediate the transport of glucose and fructose were largely upregulated in Asg and Msg of BmSUC1 mutants. In particular, transcripts associated with fatty acid metabolism exhibited obvious upregulation in the BmSUC1 mutant. In particular, the transcript levels of many genes that function in the process of fatty acid oxidation were elevated upon the absence of BmSUC1, including Fabp1, Fatp (encodes a fatty acid transporter), and Acss3 (encodes an acyl-CoA synthetase), which may compensate for the shortage of energy due to downregulated carbohydrate metabolism in the silk gland of BmSUC1 mutants. Of note, BmSUC1 mutants displayed a significant reduction in ecdysone oxidase but a large elevation in juvenile hormone-binding protein (JHBP), which could change the appropriate concentration of juvenile hormone (JH) and ecdysone in the silk glands, and affect the development of the silk gland. In addition, multiple DEGs involved in protein processing in endoplasmic reticulum were expressed at reduced levels in the mutants, including DnaJ homolog 3, heat shock protein 20.4, and heat shock protein 23.7. Moreover, we found that DEGs associated with ribosome biogenesis and protein translation and export were significantly downregulated in Msg, including large subunit ribosomal protein L3, valyl-tRNA synthetase, RNA 3′-terminal phosphate cyclase, and signal recognition particle subunit (fig. 9D and E). Thus, our data indicate that BmSUC1 is not only involved in sucrose metabolism, but also may exert broader physiological functions in the silk gland.
Discussion
HGT and gene duplication are regarded as major mechanisms that contribute to the evolutionary innovation and adaptation of organisms. The evolutionary mechanisms responsible for the retention and subsequent functional diversification of genes duplicated after HGT remain largely unexplored. Here, we provided a comprehensive evolution analysis of β-fructofuranosidase in insects. Ancient HGTs of β-fructofuranosidase have been widely found in lepidopteran species, whereas sporadic occurrences of β-fructofuranosidase were observed in coleopteran and hymenopteran species. We found that the posttransfer duplication and subsequent functional divergence of SUCs frequently occurred in lepidopteran species. In addition, we found that SUC1 is a critical sucrase in the midgut and silk gland of B. mori. These results suggest that lepidopteran β-fructofuranosidase acquired from bacteria is indispensable for a highly efficient breakdown of sucrose in the midgut and silk gland of Lepidoptera, providing significant nutritional benefits. Our experimental analyses demonstrate a direct role for β-fructofuranosidase in supporting metabolic homeostasis, which appears to be conserved throughout lepidopteran evolution. The present study underscores HGT as an important force that modulates lepidopteran evolution and niche adaptation.
We identified a number of homologs of SUC in lepidopteran species that we surveyed, indicating that SUC genes are ubiquitous in Lepidoptera. Homologous lateral gene transfers of β-fructofuranosidase have occurred sporadically in Coleoptera and Hymenoptera, raising the question of the time and number of HGT events needed to explain the phylogenetic pattern of the distribution of β-fructofuranosidase genes. It appears plausible that a single HGT event occurred within a common ancestor of these orders, followed by selective losses in different lineages, leading to the present phylogenetic distribution. However, multiple coleopteran and hymenopteran SUC-likes showed a closer phylogenetic relationship to β-fructofuranosidases of Firmicutes or Proteobacteria. Furthermore, synteny was not found near β-fructofuranosidase genes among Lepidoptera, Hymenoptera, and Coleoptera. Thus, the phylogenetic results, together with previous studies, suggest that lepidopteran, hymenopteran, and coleopteran β-fructofuranosidases have different evolutionary origins, which is against the origin of the gene in a common ancestor (Pedezzi et al. 2014; Zhao et al. 2014). SUC homologs have not been found in most examined coleopteran and hymenopteran species. Therefore, we propose that there was no occurrence of β-fructofuranosidase in the common ancestor of the three insect orders. The gain of β-fructofuranosidase genes may have occurred from multiple gene transfer events. We found that SUC genes from all examined lepidopteran species formed a monophyletic clade separated from those of the other two orders. Based on the current phylogenetic tree and SUC distribution pattern, we tentatively concluded that independent HGT events occurred, and β-fructofuranosidases in the three orders have divergent evolutionary bacteria origins. However, the current data do not allow estimations of the time of β-fructofuranosidase gene transfer among these orders.
Synteny analysis and phylogenetic tree suggest that SUC1a and SUC1b arose from the duplication of a common ancestral gene in the Lepidoptera. However, the distribution of SUC3a/3b homologs in other lepidopteran species is limited, making it difficult to establish the credibility of a tandem gene event. Hence, further studies are needed to confirm this hypothesis. Our phylogenetic analyses demonstrated that lepidopteran SUCs were separated into three clusters, and SUC1/SUC3 was grouped into two clades, separate from SUC2. Therefore, these findings suggest a high probability of a single HGT event of SUC from bacteria to a common lepidopteran ancestor, followed by gene duplication events. The presence of four SUC paralogs in five lepidopteran species (O. brumata, M. sexta, S. frugiperda, P. interpunctella, and A. transitella) suggested that duplication events occurred in their common ancestor and that copies were subsequently maintained in these species; however, one or two SUC copies may have been lost in other species. Differences in duplicated genes and copy number in these species suggest that lepidopteran SUC gene families have suffered extensive gene loss after the posttransfer duplications. Differences in the repertoires of lepidopteran SUC genes may be because of differences in the diet and living environment, which require changes in the metabolic pathways of sucrose. A previous study showed that the large number of desaturase genes present in many lepidopteran lineages reflected their enormous ecological success and diversity (Fang et al. 2009). This appears to be particularly plausible for the Lepidoptera, which colonized nearly every terrestrial habitat and are extremely diverse in terms of their diet. Therefore, we speculate that the dynamic pattern of SUC gene gain and loss in Lepidoptera may reflect an adaptation in response to ecological diversification and an increased demand for sucrose. Indeed, increasing genomic data are revealing a novel perspective of gene loss as a pervasive source of genetic variation in insects (Albalat and Cañestro 2016).
Gene duplication is a major evolutionary mechanism that can contribute genomes with the raw material for new or altered gene functions (Zhang 2003; Marques et al. 2008; Farre and Alba 2010; Baudouin-Gonzalez et al. 2017; Leite et al. 2018). When a gene is duplicated, the copies may face any of the several evolutionary fates. According to literature and empirical data, most paralogs undergo loss of function (Zhang et al. 1998; Zhang 2003; Nardelli et al. 2019). Harmful mutations occur more frequently than beneficial ones; hence, one copy may accumulate deleterious mutations, and eventually become a nonfunctional pseudogene (Baudouin-Gonzalez et al. 2017; Zhou et al. 2019). One indication of this pseudogenization is the absence of expression of the gene. In B. mori and P. xuthus, the expression of SUC2 was much weaker than that of SUC1. Three conserved active sites are essential for β-fructofuranosidase hydrolytic activity: Asp63 (D), which acts as a catalytic nucleophile; Asp181 (D), which can bind to characteristic hydroxyl groups of the substrate; and Glu234 (E), which is believed to be a proton donor to the glycosidic oxygen of sucrose (Guo, Wang, et al. 2018). These sites were not conserved in SUC2s. In addition, the recombinant SUC2 possessed no hydrolytic activity toward any of the three sugar substrates. Therefore, lepidopteran SUC2s are unlikely to be functional β-fructofuranosidases. Except in some rare cases, no deleterious mutations occur; therefore, paralogs diverge rapidly in expression pattern and/or protein function and may undergo the neofunctionalization or subfunctionalization process (Force et al. 1999; Kellogg 2003; Kondrashov and Innan 2010; Turetzek et al. 2016; Nardelli et al. 2019). We examined the expression of SUC1 and SUC3 in P. xuthus. PxSUC1 was highly expressed in the midgut and exhibited higher β-fructofuranosidase activity when sucrose was used as substrate, indicating that PxSUC1 retains a subset of functions of the ancestral gene. Unlike PxSUC1, PxSUC3 was exclusively expressed in the larval silk gland and differed in the optimal pH, suggesting that PxSUC3 gained a new adaptive function (neofunctionalization) or divided an ancestral function (subfunctionalization). Therefore, we speculate that SUC orthologs confer adaptive advantages to lepidopteran species by subfunctionalization and/or neofunctionalization.
Previous studies have indicated that the majority of lepidopteran HGT events probably function in nutrition and detoxification metabolism (Li et al. 2011; Zhu et al. 2011; Sun et al. 2013). Publicly available RNA-seq data show that the β-fructofuranosidase gene is widely present in the lepidopteran midgut, which is not only required for the digestion and absorption of nutrients but is also an essential organ for performing intermediary metabolism processes (Daimon et al. 2008; Yang et al. 2016; Gan, Zhang, et al. 2018; Guo, Wang, et al. 2018). Our transcriptome investigations revealed that the interruption of BmSUC1 elicited significant alterations in DEGs associated with carbohydrate metabolism, lipid metabolism, and amino acid metabolism, which suggests that the SUC1 acquired by HGT participates in nutritional metabolism and plays an essential role in larval metabolic homeostasis. The disruption of metabolic homeostasis may lead to the larval malnutrition and abnormal phenotypes. Based on the study results, we concluded that this HGT from bacteria to lepidopteran genomes is beneficial for organismal adaptation and evolution, thereby promoting digestive and metabolic adaptability. In addition, midgut sucrase activities were strongly inhibited by a low concentration of 1-DNJ in BmSUC1 mutants, indicating that SUC1 could promote the mulberry-specialist insect to survive on sugar-mimic alkaloid-rich diet in their ecological niches (Daimon et al. 2008). Therefore, our findings, together with previous studies, revealed that the HGT of SUC was utilized by lepidopteran mulberry-specialist insects to decrease the inhibitory effect of alkaloids on sucrose.
Lepidopteran larvae possess a pair of mandibular salivary glands, besides their labial silk glands (Afshar et al. 2013). Previous studies have shown that mandibular salivary glands are involved in digestion, and the labial silk glands become specialized for silk protein synthesis by the provision of an adequate supply of dietary carbohydrate during the final stage of larval development (Eichenseer et al. 2010). High-level expression of the β-fructofuranosidase gene was detected in the anterior and middle silk gland tissue of Lepidoptera; however, the specific function of β-fructofuranosidase in the silk gland is unclear (Daimon et al. 2008; Guo, Wang, et al. 2018). Our transcriptome analysis demonstrated that the loss of BmSUC1 induces a significant downregulation in DEGs that were functionally associated with carbohydrate metabolism. By contrast, the extensive number of DEGs encoding enzymes for fatty acid metabolism and glucose transport were largely upregulated, suggesting that these genes are involved in the compensation of nutrient and energy deficiencies due to the downregulated carbohydrate metabolism after the knockout of BmSUC1. Moreover, our results revealed that DEGs of Msg were specifically enriched in the metabolic process, oxidoreductase activity, and peptidase activity, whereas transporter activity and signal transducer were specifically enriched in Asg. This is perhaps not surprising given that sericins, as soluble glue proteins, are mainly synthesized and secreted in Msg, whereas Asg is mainly responsible for spitting out silk (Shi et al. 2019). As revealed by previous studies, JH and ecdysone regulate the development of silk gland tissues and also coordinate the synthesis of silk proteins (Guo, Zhang, et al. 2018; Liu et al. 2019). JH actions are initiated by the transport of JH in the hemolymph as a complex with JHBP to target tissues (Li et al. 2016). Our results indicated that JHBP was increased, whereas ecdysone oxidase, which converts ecdysone into 3-dehydroecdysone thus inactivating ecdysone, was downregulated in the silk gland of BmSUC1 mutants. In addition, significant DEG changes in relation to protein synthesis processes were observed. For example, six DEGs encoding heat shock proteins (HSPs) (<30 kDa) were expressed at reduced levels in Msg of BmSUC1 mutants, and two were expressed at reduced levels in Asg. A previous study has shown that HSPs act as molecular chaperones and protect cellular proteins from misfolding and aggregation (Braakman and Bulleid 2011). Moreover, we found that DEGs associated with ribosome biogenesis and protein translation and export were significantly downregulated in Msg. The absence of BmSUC1 probably indirectly leads to the downregulation of these genes. These changes probably exert a negative impact on protein synthesis, particularly in the Msg of BmSUC1 mutants. Taken together, these results support the hypothesis that the transferred BmSUC1 gene participates in the metabolic processes, and also exert broader physiological functions in the silk gland.
Conclusions
In the present study, we found that lepidopteran insects experienced specific SUC gene duplications after HGT from bacteria and multiple SUC copies were retained in their genomes. These genes have high sequence similarity but different patterns of expression and enzyme properties, implying that these genes have suffered distinct evolutionary pressures for functional divergence. To the best of our knowledge, this is the first study to comprehensively ascertain the physiological functions of β-fructofuranosidase using the CRISPR/Cas9 system in insects. Our findings provide direct evidence that the transferred BmSUC1 gene supports larvae metabolic homeostasis and also participates in carbohydrate metabolism and protein synthesis in the silk gland. Lepidopteran insects vary considerably in life-history traits, resulting in them having very different diets and being exposed to different living environments. The horizontal transmission of SUC contributed to lineage-specific adaptations regarding these differences, enabling lepidopterans to make the maximum use of sucrose from host plants. The dynamic evolution of SUC may reflect changes in carbohydrate metabolic pathways, response to ecological diversification, and an increased demand for sucrose variability. The present study highlights the importance of SUCs in coupling developmental progression with metabolism in the lepidopteran insects, which extends our knowledge about the extent and role of HGT in insects and, more broadly, eukaryotes. Taken together, based on a multilevel survey, the genomic sequence analyses of SUCs from the species level as well as functional assays and transcriptomic analyses help identify adaptive signatures that may reflect molecular adaptations to the diverse ecological challenges faced by lepidopteran species. This may contribute to a new and more complete understanding of the complex biological features of genomic evolution and organismal adaptation.
Materials and Methods
Insect Materials and Cell Lines
Papilio xuthu larvae and adults were reared at 25 °C on fresh citrus leaves. Bombyx mori larvae (Nistari) were reared on fresh mulberry leaves under standard conditions (25 °C, 12 h:12 h-l: D photoperiod and 70% relative humidity). Sf9 and High Five cells were cultured as previously described (Dai et al. 2019).
Phylogenetic Analysis
To identify the distribution and evolutionary relationships of SUC genes, BlastN and BlastP were used to search for SUC homologs in NCBI (https://www.ncbi.nlm.nih.gov/, last accessed March 21, 2021), Ensembl (http://metazoa.ensembl.org/index.html, last accessed March 21, 2021), and Lepbase databases (http://lepbase.org/, last accessed March 21, 2021) using the B. mori SUC1 and SUC2 as query sequences. Multiple amino acid sequence alignment analyses were performed using the MUSCLE program in MEGA7.0 software (v7.0.26) and were then visualized using the Geneious program (v9.1.4). ProtTest software (v3.4.2) was used to identify the optimal amino acid substitution model based on the Akaike Information Criterion for small sizes. Phylogenetic relationships were reconstructed using ML method in IQ-TREE (v2.0.6) and MEGA7.0 software. Phylogenetic trees were annotated and edited using the FigTree software (v1.4.4) and Adobe Photoshop CS6 program (v13.1.3). The alignments of three catalytic regions (62–65,180–183, and 234–237 amino acids; numbered according to BmSUC1) were extracted following an alignment of the full-length SUC protein sequences in MEGA7.0 program. The annotated amino acid sequence alignments on the phylogeny were visualized using “ggtree” in R (v3.6.2) (Yu et al. 2018). All sequences used in the construction of the phylogeny are presented in supplementary tables S1 and S4, Supplementary Material online.
Conserved Synteny Analyses
The syntenic relationship of three PxSUCs and their neighboring genes located on the same scaffold of lepidopteran genomes were examined using the Genomicus genome browser (database version 30.01) with PxSUC1, PxSUC2, and PxSUC3 as the query sequences, respectively. Some genes around the corresponding orthologs were acquired from the NCBI genome database or Lepbase database, and the result was collated via reciprocal BLAST.
Expression Analysis by Quantitative Real-Time PCR
We dissected different tissues (head, wing disc, silk gland, fat body, midgut, Malpighian tubule, epidermis, and testis) from different developmental stages (L5D3, P4, and adult) to examine the divergence of transcriptional profiles among SUC genes in P. xuthus. Quantitative real-time PCR (qRT–PCR) was used to examine the transcriptional profiles of different P. xuthu tissues from separate developmental stages (L5D3, P4, and adult). qRT–PCR assays were performed as previously described (Liu et al. 2017). The coding gene of ribosomal protein 32 (GenBank accession number NM_001312851) was used as an internal control. Relative gene expression data were analyzed using the 2−ΔΔCT method as described. All primers used for qRT–PCR are listed in supplementary table S5, Supplementary Material online. mRNA measurements were quantitated in three independent biological replicates and normalized to the gene encoding ribosomal protein (rpl).
Western Blotting
The BmSUC1 antibody was produced as previously described (Daimon et al. 2008), and the PxSUC1 antibody was produced in the Huaan biotechnology company (Hangzhou, China) as previously described (Dai et al. 2019). Sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS–PAGE) and immunoblot analyses were performed as previously described (Kiuchi et al. 2014). The anti-BmSUC1 or anti-PxSUC1 polyclonal antibody was used as the primary antibody (1:1,000), and the membrane was subsequently probed with goat antirabbit HRP-conjugated IgG (1:4,000) as the secondary antibody. Final signals were detected with FDbio-Dura ECL Kit (FD-bio science, China) using the chemiluminescence imaging system (Clinx, China).
Immunohistochemistry
Specimens of the midgut of P. xuthus larvae were embedded in paraffin wax and sectioned for immunohistochemistry. Polyclonal rabbit anti-PxSUC1 (1:200) was used as the primary antibody and a fluorescein isothiocyanate-labeled goat antirabbit immunoglobulin (1:300) (Invitrogen) was used as the secondary antibody. Nuclei were stained using 4,6-diamidino-2-phenylindole dihydrochloride solution (DAPI; Wako). The slides were then photographed under a fluorescence microscope (Nikon Nis-Elements, Japan). The preimmune serum was used a control.
Expression and Purification of Recombinant PxSUC Proteins
Three recombinant SUC proteins with a His-tag sequence were expressed as previously described (Zhou et al. 2018). Recombinant proteins were purified from the culture medium using His SpinTrap columns (GE Healthcare, Sweden) as previously described (Dai et al. 2019). Sample purity was further examined by Coomassie Brilliant Blue staining and Western-blot analysis using the antihistidine antibody. All primers used are listed in supplementary table S5, Supplementary Material online.
Enzyme Assay
The substrate specificity was assayed as previously described (Wang et al. 2015). The standard 20 μl reaction mixture contained 200 ng purified recombinant protein, 100 mM substrate (sucrose, maltose, or isomaltose), and 10 mM Britton–Robinson buffer (pH 7.0). The mixtures were incubated at 30 °C for 30 min. The reaction was stopped by heating the mixtures for 5 min at 100 °C. Glucose C II-Test Wako (Wako, Japan) was used to measure the liberated glucose at 505 nm using a spectrophotometer. The optimum pH was determined in 20 mM Britton–Robinson’s wide range buffer (pH 3.0–11.0), using sucrose as a substrate. Michaelis–Menten constants (Km and Vmax) were determined as previously described (Dai et al. 2019). All experiments were repeated in triplicate and independently conducted.
Construction of BmSUC1 Mutant, Mutation Analysis, and Phenotype Screening
CRISPR/Cas9 system was conducted as previously described (Wang et al. 2013). Cas9 mRNA was synthesized and purified using T7 MEGAscript kit (Ambion) following the manufacturer’s protocol. Two sgRNAs targeting the BmSUC1 gene were designed using ZiFit Targeter version 4.2 tool (supplementary table S5, Supplementary Material online). Customized sgRNAs were prepared as previously described with transcription performed using mMESSAGE mMACHINE T7 kit (Ambion) (Gao et al. 2020). Embryos of the Nistari strain were prepared as previously described and were injected within 6 h after oviposition. Each egg was injected with approximately 2 nl mixture containing each sgRNA (150 ng/μl) and Cas9 mRNA (300 ng/μl) (Gao et al. 2020). Injected eggs were incubated at 25 ± 1 °C and 60 ± 10% relative humidity until hatching.
To identify the targeted mutations induced by CRISPR/Cas9, we collected generation 0 (G0) moths to extract enough genomic DNA for PCR and sequencing. Mutation events were detected by amplification using specific primers designed for the upstream and downstream regions of the target site. If their male and/or female parent was tested as chimeric by sequence, the G0 moths were sibling-crossed with WT moths in single pairs. Only the sgRNA of target site 1 was highly efficient. The genotype of the G1 parent was analyzed by sequencing. G1 moths were sibling crossed with each other in single pairs to produce G2 progeny. A series of crossing strategies and persistent PCR-based multigenerational selections were performed to ensure the germline transmission of the genotype. PCR primer pairs are shown in supplementary table S5, Supplementary Material online.
To investigate the growth and development of silkworm larvae, 20 individuals from the WT or CRISPR/Cas9-mediated mutants were randomly selected from a group and fed with quantitative mulberry leaves. Each larva was fed with 3.0 g fresh mulberry leaves on the first and second days of the fifth instar and 4.0 g fresh mulberry leaves on the remaining days of the fifth instar. The weight and feeding intake of each larvae were weighed at every 12 h interval during the fifth instar stage.
Preparation of Enzyme Fractions and Inhibitor Assays
The midgut, Asg, and Msg were dissected from the larval stage and washed with phosphate buffer (pH 7.0) to remove the hemolymph and gut contents. The preparation of enzyme fractions from these tissues was performed as previously described (Hirayama et al. 2007). The BCA protein assay kit (Pierce) was utilized to determine the protein concentration in each enzyme fraction: soluble enzyme fraction, membrane-associated enzyme fraction, and membrane-bound fraction. Disaccharidase assays were performed on the three enzyme fractions of controls or BmSUC1 mutants as described above. Various concentrations of 1-DNJ (Santa Cruz Biotechnology) were used to determine the effects of the inhibitor on the enzyme activity. All experiments were performed independently in triplicate.
Library Preparation, Sequencing, and Data Processing
Total RNA was isolated from the midgut, Asg, and Msg of fifth instar larvae, and extracted using the RNA-TRIzol reagent (Invitrogen) according to the manufacturer’s protocol. Transcript library construction was performed with 1 μg of total RNA using Prime Script kit (Promega). After clustering generation, the libraries were sequenced on Illumina BGISEQ-500 platform to generate paired-end reads. The adaptor removal and data filter procedures were performed using SOAPnuke software (v1.5.2) (Cock et al. 2010). Clean reads were then mapped to the B. mori reference genome (GenBank accession number GCA_000151625.1) using Bowtie2 (v2.2.5) and HISAT2 (v2.1.0) (Langmead and Salzberg 2012; Kim et al. 2015; Kawamoto et al. 2019). The sam format file was converted into bam format using SAMtools (Li et al. 2009). The fragments per kilobase of transcript per million reads mapped (FPKM) were calculated and used to quantify the gene expression level by RSEM software (v1.2.8) (Li and Dewey 2011).
Differential Expression Analysis
Genes were represented as DEGs when false discovery rate <0.001 and |log2 (fold-change)| > 1.0. All DEGs were functionally annotated against the databases, including the NCBI nonredundant protein database (Nr), GO, and KEGG. Enrichment analysis was performed using clusterProfiler (Yu et al. 2012). In general, terms and pathways with P < 0.05 were considered as significant enrichment. Three independent hierarchies represented the results of GO enrichment: biological process, molecular function, and cellular component (supplementary tables S2 and S3, Supplementary Material online). We manually classified 43 GO terms into 11 major functional groups to study the classification features in GO terms in Asg and Msg (supplementary table S3, Supplementary Material online). The “ComplexHeatmap” of R package was utilized to visualize the differential expression analysis results (Gu et al. 2016).
Statistical Analysis
Data on the number of eggs, hatchability, weight, feeding intake, and enzyme activity were analyzed via Student’s t-test. P values <0.05 were considered statistically significant; they were represented as follows: *P < 0.05, **P < 0.01, ***P < 0.001, and n.s., P > 0.05. The data from three independent experiments were recorded and presented as means ± SEM.
Supplementary Material
Supplementary data are available at Molecular Biology and Evolution online.
Supplementary Material
Acknowledgments
We are indebted to Yan Meng, Muwang Li, and Jianhong Xu for helpful suggestions on this study. We are grateful to Boxiong Zhong, who provides the practical advices on microinjection. We thank Katsuhiko Ito, Xiaotong Li, Pu Tang, Lingyan Wang, Wenhui Jing, and Yue Jin for technical support. This work was supported by the National Natural Science Foundation of China (Grant Nos. 31970460, 31602010, and 31572321). This study was also supported, in part, by the JSPS Grants-in-Aid for Scientific Research (Grant Nos. 18H03949 and 22128004).
Author Contributions
X.D., T.K., T.S., and H.W. conceived the study, and designed the experiments. X.D., Y.Z., S.J., and H.W. performed experiments and most data analyses. T.K., Y.X., S.K., T.S., and H.W. contributed reagents or analytical tools. X.D. and H.W. wrote the manuscript draft. All authors gave final approval for publication.
Data Availability
All raw RNA-seq data are uploaded to the NCBI database (accession no. PRJNA645587) and available upon publication.
References
- Acuña R, Padilla BE, Flórez-Ramos CP, Rubio DJ, Herrera JC, Benavides P, Lee SJ, Yeats TH, Egan AN, Doyle JJ, et al. 2012. Adaptive horizontal transfer of a bacterial gene to an invasive insect pest of coffee. Proc Natl Acad Sci U S A. 109(11):4197–4202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Afshar K, Dube FF, Najafabadi HS, Bonneil E, Thibault P, Salavati R, Bede JC.. 2013. Insights into the insect salivary gland proteome: diet-associated changes in caterpillar labial salivary proteins. J Insect Physiol. 59(3):351–366. [DOI] [PubMed] [Google Scholar]
- Albalat R, Cañestro C.. 2016. Evolution by gene loss. Nat Rev Genet. 17(7):379–391. [DOI] [PubMed] [Google Scholar]
- Assis R, Bachtrog D.. 2013. Neofunctionalization of young duplicate genes in Drosophila. Proc Natl Acad Sci U S A. 110(43):17409–17414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baudouin-Gonzalez L, Santos MA, Tempesta C, Sucena É, Roch F, Tanaka K.. 2017. Diverse cis-regulatory mechanisms contribute to expression evolution of tandem gene duplicates. Mol Biol Evol. 34(12):3132–3147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Braakman I, Bulleid NJ.. 2011. Protein folding and modification in the mammalian endoplasmic reticulum. Annu Rev Biochem. 80:71–99. [DOI] [PubMed] [Google Scholar]
- Castagnone-Sereno P, Mulet K, Danchin EGJ, Koutsovoulos GD, Karaulic M, Rocha M, Marc Bailly-Bechet M, Pratx L, Perfus-Barbeoch L, Abad P.. 2019. Gene copy number variations as signatures of adaptive evolution in the parthenogenetic, plant-parasitic nematode Meloidogyne incognita. Mol Ecol. 28(10):2559–2572. [DOI] [PubMed] [Google Scholar]
- Cock PJ, Fields CJ, Goto N, Heuer ML, Rice PM.. 2010. The Sanger FASTQ file format for sequences with quality scores, and the Solexa/Illumina FASTQ variants. Nucleic Acids Res. 38(6):1767–1771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dai X, Li R, Li X, Liang Y, Gao Y, Xu Y, Shi L, Zhou Y, Wang H.. 2019. Gene duplication and subsequent functional diversification of sucrose hydrolase in Papilio xuthus. Insect Mol Biol. 28(6):862–872. [DOI] [PubMed] [Google Scholar]
- Daimon T, Taguchi T, Meng Y, Katsuma S, Mita K, Shimada T.. 2008. Beta-fructofuranosidase genes of the silkworm, Bombyx mori: insights into enzymatic adaptation of B. mori to toxic alkaloids in mulberry latex. J Biol Chem. 283(22):15271–15279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Danchin EG, Guzeeva EA, Mantelin S, Berepiki A, Jones JT.. 2016. Horizontal gene transfer from bacteria has enabled the plant-parasitic nematode Globodera pallida to feed on host-derived sucrose. Mol Biol Evol. 33(6):1571–1579. [DOI] [PubMed] [Google Scholar]
- Danchin EG, Rosso MN, Vieira P, Almeida-Engler JD, Coutinho PM, Henrissat B, Abad P.. 2010. Multiple lateral gene transfers and duplications have promoted plant parasitism ability in nematodes. Proc Natl Acad Sci U S A. 107(41):17651–17656. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dyson CJ, Goodisman MAD.. 2020. Gene duplication in the honeybee: patterns of DNA methylation, gene expression, and genomic environment. Mol Biol Evol. 37(8):2322–2331. [DOI] [PubMed] [Google Scholar]
- Eichenseer H, Mathews MC, Powell JS, Felton GW.. 2010. Survey of a salivary effector in caterpillars: glucose oxidase variation and correlation with host range. J Chem Ecol. 36(8):885–897. [DOI] [PubMed] [Google Scholar]
- Fan C, Chen Y, Long M.. 2008. Recurrent tandem gene duplication gave rise to functionally divergent genes in Drosophila. Mol Biol Evol. 25(7):1451–1458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fang S, Ting CT, Lee CR, Chu KH, Wang CC, Tsaur SC.. 2009. Molecular evolution and functional diversification of fatty acid desaturases after recurrent gene duplication in Drosophila. Mol Biol Evol. 26(7):1447–1456. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Farre D, Alba MM.. 2010. Heterogeneous patterns of gene-expression diversification in mammalian gene duplicates. Mol Biol Evol. 27(2):325–335. [DOI] [PubMed] [Google Scholar]
- Force A, Lynch M, Pickett FB, Amores A, Yan YL, Postlethwait J.. 1999. Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151(4):1531–1545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gan Q, Zhang X, Zhang D, Shi L, Zhou Y, Sun T, Jiang S, Gao J, Meng Y.. 2018. BmSUC1 is essential for glycometabolism modulation in the silkworm, Bombyx mori. Biochim Biophys Acta Gene Regul Mech. 1861(6):543–553. [DOI] [PubMed] [Google Scholar]
- Gan Q, Li X, Zhang X, Wu L, Ye C, Wang y, Gao J, Meng Y.. 2018. D181A site-mutagenesis enhances both the hydrolyzing and transfructosylating activities of BmSUC1, a novel β-fructofuranosidase in the silkworm Bombyx mori. Int J Mol Sci. 19(3):683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao Y, Liu YC, Jia SZ, Liang YT, Tang Y, Xu YS, Kawasaki H, Wang HB.. 2020. Imaginal disc growth factor maintains cuticle structure and controls melanization in the spot pattern formation of Bombyx mori. PLoS Genet. 16(9):e1008980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gogarten JP, Townsend JP.. 2005. Horizontal gene transfer, genome innovation and evolution. Nat Rev Microbiol. 3(9):679–687. [DOI] [PubMed] [Google Scholar]
- Gu Z, Eils R, Schlesner M.. 2016. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32(18):2847–2849. [DOI] [PubMed] [Google Scholar]
- Guo K, Dong Z, Zhang Y, Wang D, Tang M, Zhang X, Xia Q, Zhao P.. 2018. Improved strength of silk fibers in Bombyx mori trimolters induced by an anti-juvenile hormone compound. Biochim Biophys Acta Gen Subj. 1862(5):1148–1156. [DOI] [PubMed] [Google Scholar]
- Guo PC, Wang Q, Wang Z, Dong Z, He H, Zhao P.. 2018. Biochemical characterization and functional analysis of invertase Bmsuc1 from silkworm, Bombyx mori. Int J Biol Macromol. 107(Pt B):2334–2341. [DOI] [PubMed] [Google Scholar]
- Heidel-Fischer HM, Kirsch R, Reichelt M, Ahn SJ, Wielsch N, Baxter SW, Heckel DG, Vogel H, Kroymann J.. 2019. An insect counteradaptation against host plant defenses evolved through concerted neofunctionalization. Mol Biol Evol. 36(5):930–941. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Helmkampf M, Cash E, Gadau J.. 2015. Evolution of the insect desaturase gene family with an emphasis on social Hymenoptera. Mol Biol Evol. 32(2):456–471. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hirayama C, Konno K, Wasano N, Nakamura M.. 2007. Differential effects of sugar-mimic alkaloids in mulberry latex on sugar metabolism and disaccharidases of Eri and domesticated silkworms: enzymatic adaptation of Bombyx mori to mulberry defense. Insect Biochem Mol Biol. 37(12):1348–1358. [DOI] [PubMed] [Google Scholar]
- Hotopp JCD, Clark ME, Oliveira DCSG, Foster JM, Fischer P, Torres MCM, Giebel JD, Kumar N, Ishmael N, Wang S, et al. 2007. Widespread lateral gene transfer from intracellular bacteria to multicellular eukaryotes. Science 317(5845):1753–1756. [DOI] [PubMed] [Google Scholar]
- Husnik F, McCutcheon JP.. 2018. Functional horizontal gene transfer from bacteria to eukaryotes. Nat Rev Microbiol. 16(2):67–79. [DOI] [PubMed] [Google Scholar]
- Kawamoto M, Jouraku A, Toyoda A, Yokoi K, Minakuchi Y, Katsuma S, Fujiyama A, Kiuchi T, Yamamoto K, Shimada T.. 2019. High-quality genome assembly of the silkworm, Bombyx mori. Insect Biochem Mol Biol. 107:53–62. [DOI] [PubMed] [Google Scholar]
- Keeling CI, Yuen MM, Liao NY, Roderick Docking T, Chan SK, Taylor GA, Palmquist DL, Jackman SD, Nguyen A, Li M, et al. 2013. Draft genome of the mountain pine beetle, Dendroctonus ponderosae Hopkins, a major forest pest. Genome Biol. 14(3):R27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keeling PJ. 2009. Functional and ecological impacts of horizontal gene transfer in eukaryotes. Curr Opin Genet Dev. 19(6):613–619. [DOI] [PubMed] [Google Scholar]
- Keeling PJ, Palmer JD.. 2008. Horizontal gene transfer in eukaryotic evolution. Nat Rev Genet. 9(8):605–618. [DOI] [PubMed] [Google Scholar]
- Kellogg EA. 2003. What happens to genes in duplicated genomes. Proc Natl Acad Sci U S A. 100(8):4369–4371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim D, Langmead B, Salzberg SL.. 2015. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 12(4):357–360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kiuchi T, Koga H, Kawamoto M, Shoji K, Sakai H, Arai Y, Ishihara G, Kawaoka S, Sugano S, Shimada T, et al. 2014. A single female-specific piRNA is the primary determiner of sex in the silkworm. Nature 509(7502):633–636. [DOI] [PubMed] [Google Scholar]
- Kondrashov F, Innan H.. 2010. The evolution of gene duplications: classifying and distinguishing between models. Nat Rev Genet. 11(2):97–108. [DOI] [PubMed] [Google Scholar]
- Konno K, Ono H, Nakamura M, Tateishi K, Hirayama C, Tamura Y, Hattori M, Koyama A, Kohno K.. 2006. Mulberry latex rich in antidiabetic sugar-mimic alkaloids forces dieting on caterpillars. Proc Natl Acad Sci U S A. 103(5):1337–1341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kursel LE, Malik HS.. 2017. Recurrent gene duplication leads to diverse repertoires of centromeric histones in Drosophila species. Mol Biol Evol. 34(6):1445–1462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Langmead B, Salzberg SL.. 2012. Fast gapped-read alignment with Bowtie 2. Nat Methods. 9(4):357–359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee BH, Eskandari R, Jones K, Reddy KR, Calvillo RQ, Nichols BL, Rose DR, Hamaker BR, Pinto BM.. 2012. Modulation of starch digestion for slow glucose release through “toggling” of activities of mucosal α-glucosidases. J Biol Chem. 287(38):31929–31938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leite DJ, Baudouin-Gonzalez L, Iwasaki-Yokozawa S, Lozano-Fernandez J, Turetzek N, Akiyama-Oda Y, Prpic NM, Pisani D, Oda H, Sharma PP, et al. 2018. Homeobox gene duplication and divergence in arachnids. Mol Biol Evol. 35(9):2240–2253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li B, Dewey CN.. 2011. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12(12):323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R, 1000 Genome Project Data Processing Subgroup. 2009. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25(16):2078–2079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li W, Cheng T, Hu W, Peng Z, Liu C, Xia Q.. 2016. Genome-wide identification and analysis of JHBP-domain family members in the silkworm Bombyx mori. Mol Genet Genomics. 291(6):2159–2171. [DOI] [PubMed] [Google Scholar]
- Li X, Shi L, Dai X, Chen Y, Xie H, Feng M, Chen Y, Wang H.. 2018. Expression plasticity and evolutionary changes extensively shape the sugar-mimic alkaloid adaptation of nondigestive glucosidase in lepidopteran mulberry-specialist insects. Mol Ecol. 27(13):2858–2870. [DOI] [PubMed] [Google Scholar]
- Li X, Shi L, Zhou Y, Xie H, Dai X, Li R, Chen Y, Wang H.. 2017. Molecular evolutionary mechanisms driving functional diversification of α-glucosidase in Lepidoptera. Sci Rep. 7:45787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li ZW, Shen YH, Xiang ZH, Zhang Z.. 2011. Pathogen-origin horizontally transferred genes contribute to the evolution of lepidopteran insects. BMC Evol Biol. 11:356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu L, Wang Y, Li Y, Ding C, Zhao P, Xia Q, He H.. 2019. Cross-talk between juvenile hormone and ecdysone regulates transcription of fibroin modulator binding protein-1 in Bombyx mori. Int J Biol Macromol. 128:28–39. [DOI] [PubMed] [Google Scholar]
- Liu Y, Su H, Li R, Li X, Xu Y, Dai X, Zhou Y, Wang H.. 2017. Comparative transcriptome analysis of Glyphodes pyloalis Walker (Lepidoptera: Pyralidae) reveals novel insights into heat stress tolerance in insects. BMC Genomics 18(1):974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marques AC, Vinckenbosch N, Brawand D, Kaessmann H.. 2008. Functional diversification of duplicate genes through subcellular adaptation of encoded proteins. Genome Biol. 9(3):R54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nardelli A, Vecchi M, Mandrioli M, Manicardi GC.. 2019. The evolutionary history and functional divergence of trehalase (treh) genes in insects. Front Physiol. 10:62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pedezzi R, Fonseca FP, Santos Júnior CD, Kishi LT, Terra WR, Henrique-Silva F.. 2014. A novel β-fructofuranosidase in Coleoptera: characterization of a β-fructofuranosidase from the sugarcane weevil, Sphenophorus levis. Insect Biochem Mol Biol. 55:31–38. [DOI] [PubMed] [Google Scholar]
- Ren FR, Sun X, Wang TY, Yao YL, Huang YZ, Zhang X, Luan JB.. 2020. Biotin provisioning by horizontally transferred genes from bacteria confers animal fitness benefits. ISME J. 14(10):1–12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi R, Ma S, He T, Peng J, Zhang T, Chen X, Wang X, Chang J, Xia Q, Zhao P, et al. 2019. Deep insight into the transcriptome of the single silk gland of Bombyx mori. Int J Mol Sci. 20(10):2491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soucy SM, Huang J, Gogarten JP.. 2015. Horizontal gene transfer: building the web of life. Nat Rev Genet. 16(8):472–482. [DOI] [PubMed] [Google Scholar]
- Sun BF, Xiao JH, He SM, Liu L, Murphy RW, Huang DW.. 2013. Multiple ancient horizontal gene transfers and duplications in lepidopteran species. Insect Mol Biol. 22(1):72–87. [DOI] [PubMed] [Google Scholar]
- Turetzek N, Pechmann M, Schomburg C, Schneider J, Prpic NM.. 2016. Neofunctionalization of a duplicate dachshund gene underlies the evolution of a novel leg segment in arachnids. Mol Biol Evol. 33(1):109–121. [DOI] [PubMed] [Google Scholar]
- Wang B, Wangkahart E, Secombes CJ, Wang T.. 2019. Insights into the evolution of the suppressors of cytokine signaling (SOCS) gene family in vertebrates. Mol Biol Evol. 36(2):393–411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang H, Kiuchi T, Katsuma S, Shimada T.. 2015. A novel sucrose hydrolase from the bombycoid silkworms Bombyx mori, Trilocha varians, and Samia cynthia ricini with a substrate specificity for sucrose. Insect Biochem Mol Biol. 61:46–52. [DOI] [PubMed] [Google Scholar]
- Wang H, Sun S, Ge W, Zhao L, Hou B, Wang K, Lyu Z, Chen L, Xu S, Guo J, et al. 2020. Horizontal gene transfer of Fhb7 from fungus underlies Fusarium head blight resistance in wheat. Science 368(6493):eaba5435. [DOI] [PubMed] [Google Scholar]
- Wang Y, Li Z, Xu J, Zeng B, Ling L, You L, Chen Y, Huang Y, Tan A.. 2013. The CRISPR/Cas system mediates efficient genome engineering in Bombyx mori. Cell Res. 23(12):1414–1416. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wheeler D, Redding AJ, Werren JH.. 2013. Characterization of an ancient lepidopteran lateral gene transfer. PLoS One 8(3):e59262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wybouw N, Pauchet Y, Heckel DG, Van Leeuwen T.. 2016. Horizontal gene transfer contributes to the evolution of arthropod herbivory. Genome Biol Evol. 8(6):1785–1801. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xia Q, Zhou Z, Lu C, Cheng D, Dai F, Li B, Zhao P, Zha X, Cheng T, Chai C, Biology Analysis Group, et al. 2004. A draft sequence for the genome of the domesticated silkworm (Bombyx mori). Science 306(5703):1937–1940. [DOI] [PubMed] [Google Scholar]
- Yang B, Huang W, Zhang J, Xu Q, Zhu S, Zhang Q, Beerntsen BT, Song H, Ling E.. 2016. Analysis of gene expression in the midgut of Bombyx mori during the larval molting stage. BMC Genomics 17(1):866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu G, Lam TTY, Zhu H, Guan Y.. 2018. Two methods for mapping and visualizing associated data on phylogeny using ggtree. Mol Biol Evol. 35(12):3041–3043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu G, Wang LG, Han Y, He QY.. 2012. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16(5):284–287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang J. 2003. Evolution by gene duplication: an update. Trends Ecol Evol. 18(6):292–298. [Google Scholar]
- Zhang J, Rosenberg HF, Nei M.. 1998. Positive Darwinian selection after gene duplication in primate ribonuclease genes. Proc Natl Acad Sci U S A. 95(7):3708–3713. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao C, Doucet D, Mittapalli O.. 2014. Characterization of horizontally transferred β-fructofuranosidase (ScrB) genes in Agrilus planipennis. Insect Mol Biol. 23(6):821–832. [DOI] [PubMed] [Google Scholar]
- Zhou Y, Li X, Katsuma S, Xu Y, Shi L, Shimada T, Wang H.. 2019. Duplication and diversification of trehalase confers evolutionary advantages on lepidopteran insects. Mol Ecol. 28(24):5282–5298. [DOI] [PubMed] [Google Scholar]
- Zhou Y, Wang L, Li R, Liu M, Li X, Su H, Xu Y, Wang H.. 2018. Secreted glycoprotein BmApoD1 plays a critical role in anti-oxidation and anti-apoptosis in Bombyx mori. Biochem Biophys Res Commun. 495(1):839–845. [DOI] [PubMed] [Google Scholar]
- Zhu B, Lou M, Xie G, Zhang G, Zhou X, Li B, Jin G.. 2011. Horizontal gene transfer in silkworm, Bombyx mori. BMC Genomics 12:248. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All raw RNA-seq data are uploaded to the NCBI database (accession no. PRJNA645587) and available upon publication.