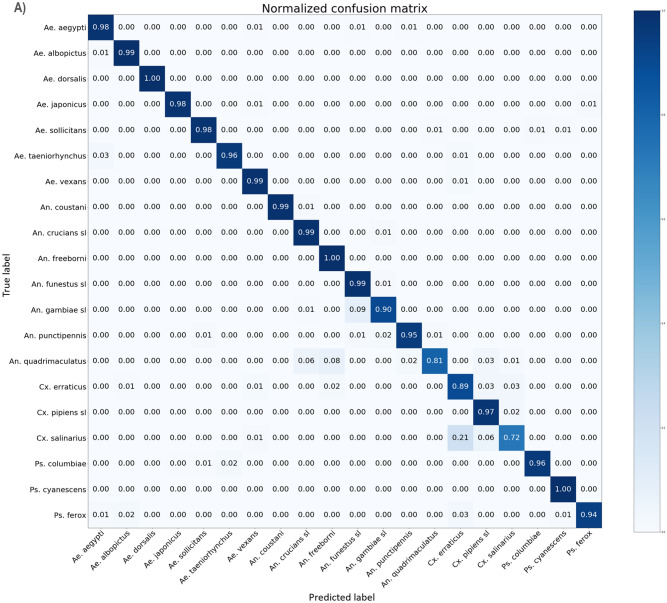
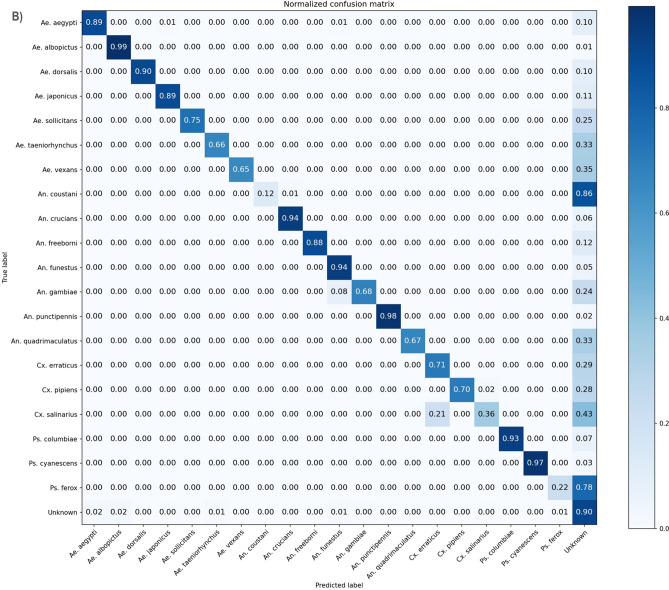
Figure 3.
Mean normalized confusion matrices for species classification shows the distribution of error within species. The species classification in these confusion matrices was performed by the Tier I CNN, the closed-set Xception model. The confusion matrix conveys the ground truth of the sample horizontally, labels on the left, and the prediction of the full methods vertically, labels on the bottom. Accurate classification is across the diagonal, where ground truth and prediction match, and all other cells on the matrix describe the error. Sixteen species were known for a given fold, and 51 species were considered unknown for a given fold, with each of the twenty known species considered unknown for one fold. (A) The species classification independent of novelty detection shows an average accuracy of 97.04 ± 0.87% and a macro F1-score of 96.64 ± 0.96%, calculated over the five folds of Tier I classifiers, trained and tested over an average of 7174.8 and 1544.6 samples. Of the error, 73.5% occurred with species of the same genus as the true species. (B) The species classification as a subsequent step after novelty detection yielded 89.07 ± 5.58% average accuracy, and a macro F1-score of 79.74 ± 3.65% trained and tested on an average of 7174.8 and 519.44 samples, evaluated over the twenty-five folds of the novelty detection methods. First, a sample was sent to the novelty detection algorithm. If the sample was predicted to be known to the species classifier, which was the closed-set Xception algorithm used in Tier I, then the sample was sent to the algorithm for classification.