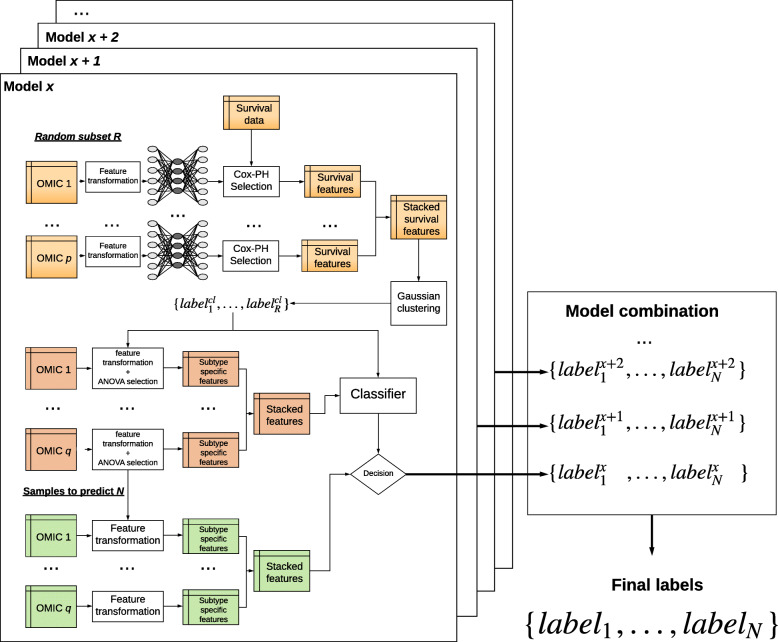
Fig. 1.
The computational framework of DeepProg. DeepProg uses the boosting strategy to build several models from a random subset of the dataset. For each model, each omic data matrix is normalized and then transformed using an autoencoder. Each of the new hidden-layer features in autoencoder is then tested for association with survival using univariate Cox-PH models. The features significantly associated with survival are then subject to clustering (Gaussian clustering by default). Upon determining the optimal cluster, the top features in each omic input data type are selected through Kruskal-Wallis analysis (default threshold = 0.05). Finally, these top omics features are used to construct a support vector machine (SVM) classifier and to predict the survival risk group of a new sample. DeepProg combines the outputs of all the classifier models to produce more robust results