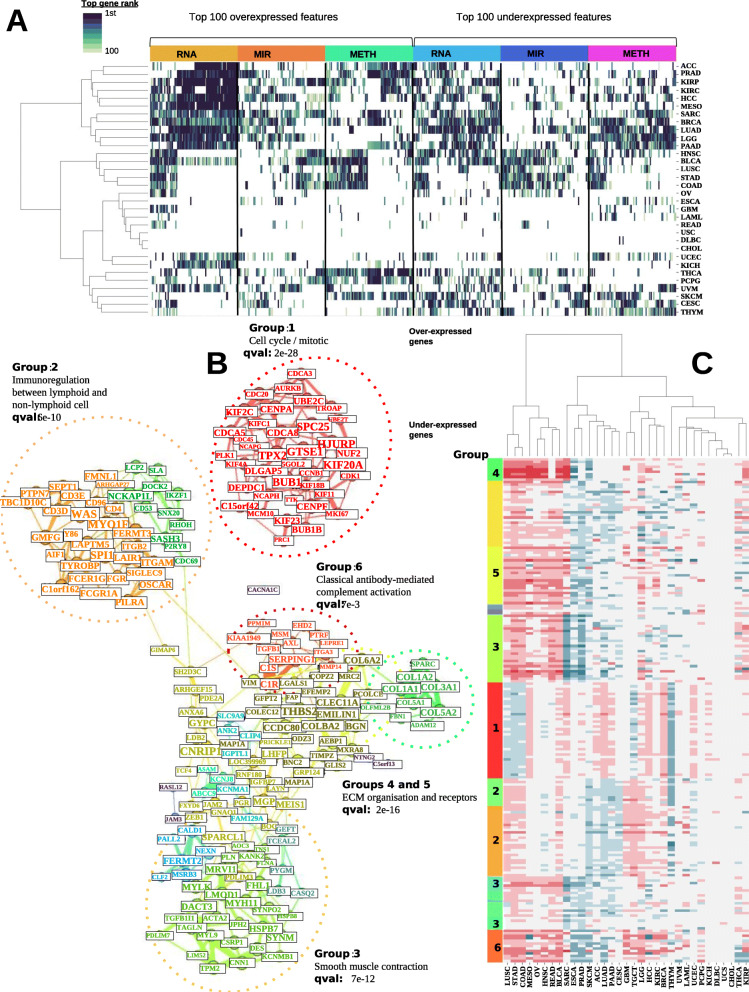
Fig. 5.
Pan-cancer analysis of RNA-Seq gene signatures in the worst survival vs. other groups. A Top 100 over- and under-expressed genes for RNA, MIR, and METH omics ranked by survival predictive power. The colors correspond to the ranks of the genes based on their –log10 (log-rank p value) of the univariate Cox-PH model. Based on these scores, the 32 cancers and the features are clustered using the WARD method. B Co-expression network constructed with the top 200 differentially expressed genes from the 32 cancers. The 200 genes are clustered from the network topology with the Louvain algorithm. For each submodule, we identified the most significantly enriched pathway as shown on the figure. C The expression values of these 200 genes used to construct the co-expression network. A clustering of the cancers using these features with the WARD method is represented in the x-axis