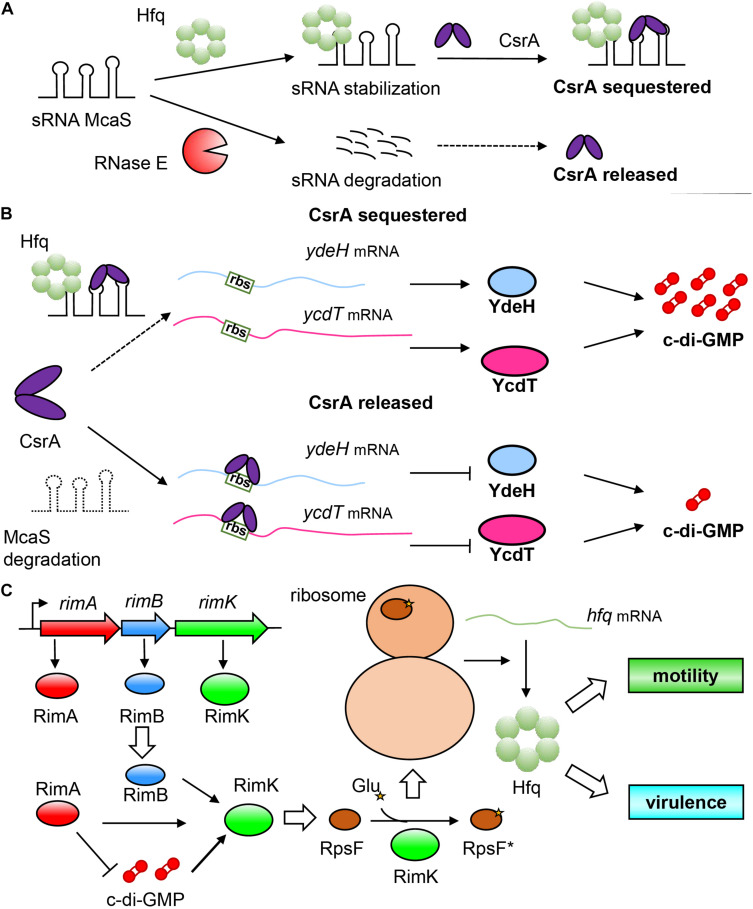
FIGURE 4.
Indirect and mutual regulation between Hfq and c-di-GMP systems. (A) A complicated regulatory relationship between sRNA McaS and RNA binding proteins Hfq and CsrA. sRNA McaS can bind to two RNA binding proteins CsrA and Hfq through different binding motifs. When Hfq is present, McaS is stabilized to adopt a conformation non-degradable by RNase E. The stable McaS then binds with CsrA, disabling its regulation on target mRNA; on the contrary, when Hfq is not present, McaS will be degraded, allowing free CsrA to play a regulatory role on target mRNA. (B) Hfq regulates the concentration of c-di-GMP by affecting the stability of sRNA McaS, thereby causing McaS to capture CsrA. In the presence of Hfq, McaS adopts a stable conformation, and binds strongly with CsrA to prevent it from blocking the ribosome binding sites of ydeH and ycdT mRNA, so that they can be translated normally (The above is reasonable speculation); while without Hfq, McaS cannot adopt a stable conformation to capture CsrA, which is then released to bind to the ribosome binding site of ydeH and ycdT mRNA, resulting in translation inhibition. (C) c-di-GMP and RimA, RimB, RimK, and RpsF co-regulate regulate Hfq. RimA (a PDE), RimB and RimK are encoded by the operon rimABK. Importantly, RimK is a glutamate ligase, which can transfer glutamate residues to the C-terminus of RpsF (Protein S6), so that RpsF is activated to become RpsF* (asterisks indicate the glutamate residues), which can interact with ribosomal proteins to regulate downstream gene activity and fine-tune the entire proteome of bacteria, thereby affecting the expression of Hfq protein.