Highlights
-
•
Proteomics of healthy mouse IVDs differentiating compartments and spine levels.
-
•
NP cells feature vacuoles with lysosomal, transport and cell–cell communication functions.
-
•
Collagen XII, decorin and other ECM proteins contribute to function of the AF.
-
•
Distinct proteomics between lumbar and tail discs.
-
•
Mouse is a relevant model for human disc biology but care is needed in its use.
Abbreviations: AF, annulus fibrosus; DEP, differentially expressed protein; ECM, extracellular matrix; IAF, inner AF; IDD, intervertebral disc degeneration; IVD, intervertebral disc; NP, nucleus pulposus; NLC, notochordal-like cell; OAF, outer AF
Keywords: Mouse intervertebral disc, Label free quantitation, Proteomics, Extracellular matrix, Nucleus pulposus, Annulus fibrosus
Abstract
Mice are commonly used to study intervertebral disc (IVD) biology and related diseases such as IVD degeneration. Discs from both the lumbar and tail regions are used. However, little is known about compartmental characteristics in the different regions, nor their relevance to the human setting, where a functional IVD unit depends on a homeostatic proteome. Here, we address these major gaps through comprehensive proteomic profiling and in-depth analyses of 8-week-old healthy murine discs, followed by comparisons with human. Leveraging on a dataset of over 2,700 proteins from 31 proteomic profiles, we identified key molecular and cellular differences between disc compartments and spine levels, but not gender. The nucleus pulposus (NP) and annulus fibrosus (AF) compartments differ the most, both in matrisome and cellularity contents. Differences in the matrisome are consistent with the fibrous nature required for tensile strength in the AF and hydration property in the NP. Novel findings for the NP cells included an enrichment in cell junction proteins for cell–cell communication (Cdh2, Dsp and Gja1) and osmoregulation (Slc12a2 and Wnk1). In NP cells, we detected heterogeneity of vacuolar organelles; where about half have potential lysosomal function (Vamp3, Copb2, Lamp1/2, Lamtor1), some contain lipid droplets and others with undefined contents. The AF is enriched in proteins for the oxidative stress responses (Sod3 and Clu). Interestingly, mitochondrial proteins are elevated in the lumbar than tail IVDs that may reflect differences in metabolic requirement. Relative to the human, cellular and structural information are conserved for the AF. Even though the NP is more divergent between mouse and human, there are similarities at the level of cell biology. Further, common cross-species markers were identified for both NP (KRT8/19, CD109) and AF (COL12A1). Overall, mouse is a relevant model to study IVD biology, and an understanding of the limitation will facilitate research planning and data interpretation, maximizing the translation of research findings to human IVDs.
Introduction
The intervertebral disc (IVD) is a cartilaginous joint situated between vertebral bodies, providing support and flexibility to the spine [1]. Human IVD degeneration (IDD) is a prevalent condition associated with aging, characterized by impaired and dysfunctional structures, that can cause severe back pain, a major disability burden with huge socioeconomic impacts [2]. It is projected to worsen with the increasing global ageing population [3]. Despite rapid progress in studying the IVD and its degeneration, treatments are limited due to major gaps in current understanding of its biology, and relevant animal models to study IVD biology, degeneration and repair that are applicable to human.
The human IVDs are composed of distinct components: the annulus fibrosus (AF), nucleus pulposus (NP) and cartilage endplates that work collectively to ensure proper function [4] which absorbs and cushions loading from daily activities. The hydrated NP is encased between cartilage endplates of adjacent vertebral bodies and confined within the concentric AF lamellae. They function together to resist the high swelling pressure of the NP, and the circumferential and tensile forces of the AF allows bending or twisting [5]. Functionality of the IVDs is facilitated by characteristic extracellular matrix (ECM) components in each compartments [1]. In health, the NP is well hydrated from an abundance of proteoglycans [6]. However, with degeneration, loss of proteoglycans and changes in other ECM reduces the hydration property, altering IVD biomechanics, affecting cellular function and IVD homeostasis.
Comprehending the changes in the ECM with aging and degeneration is key to further our understanding of disc biology that can lead to biological therapies. The most direct way to study the ECM is via proteomics as it is accumulated and modified over time. This has been demonstrated in multiple human studies by us [7], [8], and others [9], [10], [11], [12]. Recently, we presented a comprehensive proteomic atlas of the human IVDs (DIPPER) that revealed novel spatiotemporal dynamics during homeostasis, ageing and degeneration [7]. The human studies provided key insights on IVD biology directly relevant to degeneration. Nonetheless, there are challenges in studies using human tissues, such as small sample sizes, fragmented tissue sampling of surgical specimens, age matching, and difficulties in performing cell lineage studies. Further, tissues from IDD patients are of end-stage degeneration that lack information on disease progression. Animal models including bovine [13], porcine [14], canine [15] and murine [16], [17], [18], have been used to study IVD development and degeneration, each with their own advantages and limitations from variations in size, structure and cellularity [19]. Murine models, despite the size and biomechanical differences, offer a number of advantages such as genetic consistency of inbred strains, short breeding cycle and age span, and numerous degeneration models [20], providing opportunities to study gene function and assessment of cell biology using various genetic tools [21].
Anatomic and cellular differences between human and mouse IVDs are well noted, such as biomechanical loading, size and cellularity. In human, no significant differences in the prevalence of IDD between gender was observed [22]; however, some studies reported more severe disc narrowing in females [23], [24]. Further, degeneration is more prevalent at the lumbar region, attributed to the weight-bearing dynamics of the spine [25]. However, whether these and other characteristics of human IDD can be modeled in the various animal models remains unclear, limiting the translational values of the findings to the human condition. Given the mouse is commonly used to study spine biology and IVD changes, it would be important to ascertain its relevance. Therefore, a benchmark of the molecular makeup at high resolution is needed for young and healthy mouse IVDs at relevant levels of the spine, for meaningful assessment and comparison with human IVDs.
Here, we undertook a systematic and comprehensive proteomic profiling of “young adult” mouse IVDs, to establish the characteristics of “healthy” discs and generation of proteomic dataset PRIMUS (Proteomic Resource for Intervertebral disc of MUS musculus) that complements the human DIPPER dataset [7]. We performed label-free quantitative proteomics on 31 biological replicates, assessing differences between compartments (NP versus AF), spine levels (lumbar versus tail), and gender. Differentially expressed proteins (DEPs) across these phenotypic factors were determined. Pathway analyses identified top enriched functional cellular processes, and the extracellular matrix environment. Key findings were validated, providing spatial information on the marker proteins. Importantly, we cross referenced the proteome with available mouse AF and NP transcriptome datasets and assessed the relevance of the findings to young healthy human IVDs. This study addresses the applicability of using mouse models for understanding disc biology and its relevance to the human condition.
Results
Quality proteomic data detects high numbers of core and non-core matrisome proteins
To achieve a comprehensive characterization of a “healthy” mouse IVD, we performed label-free deep proteomics (by LC-MS/MS; Methods and Supplementary materials) on NP and AF of 8-week-old Foxa2mNE-Cre/ZEG mice (on mixed C57BL/6J and B6CBAF1/J backgrounds) [26], in which the NP are tagged by enhanced green fluorescent protein (eGFP). This was followed by systemic analyses and validation (Fig. 1A). We sacrificed five litters of mice, a total of 32 animals. For each litter, each of the four tissues (LAF, LNP, TAF, and TNP) (Fig. 1A) were harvested and littermates of the same gender were pooled to a total of 40 samples (5 litters × 4 tissue types × 2 gender). Nine samples did not yield sufficient materials for mass-spectrometry (Supplemental Fig. 1A). Thus, a dataset consisting of 31 biological samples was obtained, with each sample derived from a single animal or pooled from up to 5 animals (median 3.5) (Supplemental Fig. 1B).
Fig. 1.
Outline of the samples and workflow, and overview of the data. (A) Schematic depicting the samples used, proteomic workflow and type of data comparisons. (B) Barchart showing the numbers of proteins detected in each of the 31 biological replicates, categorized into matrisome (coloured) or non-matrisome proteins (grey). (C) Principal component analysis (PCA) of the 31 biological profiles. Scatter-plot of PC1 and PC2 (top) and PC1 and PC3 (bottom) color-coded by compartments and levels, and dots shaped by gender. Solid curves are the support vector machines (SVMs) decision boundaries the compartments or levels, and dashed curves are soft boundaries for probability equal to ±0.5.
To ascertain and minimize technical variation, all except one sample were measured in technical triplicates (total 91; Supplemental Fig. 1A). The MS/MS data were processed by MaxQuant [27], which detected a total of 506,507 valid MS/MS peptide-spectra matches across all IVD samples, corresponding to 25,424 unique peptide sequences and 2,731 unique proteins. Good technical fidelity was achieved, with Pearson correlation coefficients among technical replicates ranging 0.98–0.99 (Supplemental Fig. 1C). The technical triplicates were then averaged for each biological replicate, each considered as one sample (total 31) for subsequent comparative analyses.
On average, 659 proteins (min 117, max 1707, median 581; Fig. 1B) were detected per sample. An average of 17.9 % of the proteins detected in each sample were ‘matrisome’ proteins (ECM or ECM-associated proteins [28]). The fractions of matrisome proteins per sample had a negative linear correlation with increasing numbers of total proteins detected (Supplemental Fig. 1D), which could be due to the higher variability of the “non-matrisome proteins” or technical variables such as cross-samples differences in cellular contents (Fig. 1B).
Among the 2731 detected proteins, 240 were ‘matrisome’ proteins and 2491 were non-matrisome proteins (Supplemental Fig. 1E). Similar numbers of matrisome proteins (AF, n = 196; NP, n = 173) were detected in the AF and NP. Relatively, there were more ‘core matrisome’ proteins detected in the AF; whereas in the NP there were more ‘non-core matrisome’ (or ‘ECM-associated’) proteins. Of the 888 non-matrisome proteins in AF, 727 were common with the NP (Supplemental Fig. 1E), while NP had more than double the number of non-matrisome proteins than AF, likely to be related to differences in cell number and/or cellular activities. In all, the data are technically reliable and biologically informative. The complete proteomics data set is presented in Supplemental Table 1.
Proteomic profile captured differences between IVD compartments and disc levels
To gain an unbiased overview of the sample-sample relationships in the data, we performed principal component analyses (PCA) on a set of informative genes followed by imputations (see Methods). The first two principal components (PCs) explained a combined 56.8% of total variance, with 37.6 % and 19.2 % by the first and second PCs, respectively (Fig. 1C). A support vector machine (SVM) with polynomial kernel was trained to predict the boundaries between different groups of samples. The results showed PC1 can readily differentiates the NP and AF compartments (Fig. 1C, vertical boundary), whereas PC2 can distinguish the tail and lumbar IVD levels (Fig. 1C, horizontal boundary). PC3 captured 10.3 % of the variance (Fig. 1C, bottom row), with no obvious association with any of the three “phenotypes”.
AF and NP have different matrisome compositions
Next, we determined the relative proportion of proteins that defines the differences between disc compartments, IVD levels (lumbar vs tail) and gender, for core (collagens, proteoglycans and glycoproteins), non-core (ECM affiliated, ECM regulators and secreted factors) matrisome, and non-matrisome proteins [28]. When we assessed core, non-core and non-matrisome proteins as whole IVD (AF + NP), there were no differences in protein numbers between disc levels and gender (Fig. 2A). However, a significant difference was observed between AF and NP for the number of core matrisome (AF median of 66.5, NP median of 44) and non-matrisome proteins (AF median of 324, NP median of 829); but not for non-core matrisome proteins (Fig. 2A). In particular, numbers of core matrisome subcategories (collagens, proteoglycans and glycoproteins) were consistently higher in AF than NP (Supplemental Fig. S2A), as opposed to non-core matrisome subtypes (Supplemental Fig. S2A). This AF and NP difference was consistent when we compared tail and lumbar separately (Fig. 2A, Supplemental Fig. S2A).
Fig. 2.
The composition and distribution of detected proteins and their implications for cellularity. (A) Violin plots showing numbers of matrisome and non-matrisome proteins detected across compartment, level and gender groups. Red numbers on the top indicate the median. (B) Violin plots of the protein expression levels (log2LFQ) of matrisome and non-matrisome proteins across the compartment, level and gender groups. Red numbers on the top indicate median values. (C) Barchart showing the percentage of variance as explained by the compartments, levels and gender, for data encompassing the matrisome proteins, its sub-categories and the non-matrisome proteins. (D) Barchart showing the gene families for all the non-matrisome proteins detected in the dataset. (E) Scatter-plot showing the co-linearity among the histones. (F) Violin plot of the expression level (log2LFQ) of histones (top) and Gapdh (bottom) of NP and AF compartments. (For interpretation of the references to color in this figure legend, the reader is referred to the web version of this article.)
The number of non-matrisome proteins detected was higher than matrisome proteins (Fig. 1B); however, their expression levels (in terms of ‘label-free quantitation’, or LFQ values) were lower, and thus their relative abundance was also lower by comparison (Fig. 2B), consistent with IVD as a matrisome-enriched tissue. When assessed as a whole disc (AF + NP), there were no statistical differences between gender and between disc levels (Fig. 2B) for core matrisome; however, AF had significantly higher expression levels than NP that was also observed when tail and lumbar were compared separately (Fig. 2B). These were significantly attributed by all subcategories of core-matrisome (collagens, proteoglycans and glycoproteins; Supplemental Fig. S2B). A heatmap showing the expression of all the 240 matrisome proteins detected across all 31 biological replicates is shown in Supplemental Fig. S3A.
To further delineate how compartment, level and gender impacted on the protein profiles, ANOVA (analysis of variance) was performed for each of these factors for the different matrisome and non-matrisome categories of proteins (Fig. 2C; Supplemental Fig. S4). Again, the major factor that explained the highest degree of variance for both matrisome and non-matrisome proteins was compartment (Fig. 2C; Supplemental Fig. S4), consistent across all comparisons. Interestingly, disc levels (lumbar/tail) explained about 10 % of the variance across the matrisome proteins, except for proteoglycans (1.2 %). Gender with around 1 % of the data variance showed little or no impact on the protein profiles (Fig. 2C; Supplemental Fig. S4). Overall, the different assessments indicate distinctive protein compositions can be attributed to NP and AF and to a lesser degree, by disc levels.
Given there appears to be a lower level of separation with gender in the PCA (Fig. 1C), we investigated this for potential gender specific DEPs. Consequently, no DEPs were detected between male (n = 14) and female (n = 17) IVDs (Supplemental Fig. S5C & D). Likewise, no gender-related DEPs were identified when partitioned into sub-compartments (LNP, LAF, TNP or TAF) (Supplemental Fig. S5E-F). We conclude that there are no gender differences within the current dataset, and male and female samples were combined in all subsequent studies to enhance statistical power.
Histone content informs cellularity differences between NP and AF in the mouse IVD
We first asked how the functional categories of the proteins detected reflect compartmental differences. Over 80 % of the proteins detected per sample or in aggregate were non-matrisome proteins. Categorizing them into gene families revealed top major intracellular processes including RNA binding proteins, proteasome and mitochondrial function (Fig. 2D). Furthermore, we detected 69 transcription factors/DNA binding proteins and 42 cell surface markers that clustered into heatmaps (Supplemental Fig. S3B-C). There were five subunits of histone proteins expressed in most samples (Supplemental Fig. S3B), with co-expression that are highly correlative (Fig. 2E). As histones are ‘rulers’ of cellularity in tissues [29], we confirmed that the NPs have consistently higher expression levels than the AF (Fig. 1G). This together with a similar correlation with housekeeping gene, Gapdh (Fig. 1G), support a higher cell content in the mouse NP than the AF, consistent with histological findings of the mouse IVD [30], which is in contrast to lower NP cell density reported in the human IVDs [7].
Dominant expression of intracellular proteins in NP whilst ECM in AF reflects their specialized functions
To tease out the specific compartmental differences, we identified DEPs between the AF and NP, as PCA and ANOVA revealed it to be the dominant factor in explaining the overall data variability (Fig. 1C, 2C). Among the 864 proteins expressed that are common in the AF and NP samples (Fig. 3B), we detected 88 DEPs, of which 47 were higher in the NP (blue) and 41 higher in the AF (red), as shown on a volcano plot (Fig. 3A). DEPs higher in the NP included known notochordal markers such as Krt8, Krt19 and Ca3 [15], [31], [32], [33], in addition to Cd109 (an antagonist of the TGFβ pathway [34]), cytoskeletal proteins (Vim, Tuba1b, Tubb2a/4b/5, Actn4, Flnb, Sptbn1), heat shock proteins and chaperones (HSp90ab1, Hspb1, Tcp1) and lysosomal proteins (Lamp2, Atp6v1b2) (Fig. 3A), indicative of a dominance of intracellular processes in NP.
Fig. 3.
Differentially expressed proteins (DEPs) in compartments and levels. (A) Volcano plot showing the DEPs between all NP and all AF. Coloured dots represent statistically significant DEPs (FDR q < 0.05). (B) Venn diagram showing the proteins detected in NP, AF or both. (C) Heatmap showing the proteins exclusively detected in NP or AF. (D) Volcano plot showing the DEPs between the lumbar and tail discs. Coloured dots represent statistically significant DEPs (FDR q < 0.05). (E) Venn diagram showing the proteins detected in lumbar, tail or both. (F) Heatmap showing the proteins exclusively detected in lumbar or tail. (G) Venn diagram of proteins that were higher in lumbar samples (versus all tail samples), those higher in lumbar NP (versus all tail NP) and those higher in lumbar AF (versus all tail AF). (H) Venn diagram of proteins that were higher in tail samples (versus all lumbar samples), those higher in tail NP (versus lumbar NP) and those higher in tail AF (versus all lumbar AF). N represents the numbers of DEPs.
By comparison, DEPs in the AF were dominated by ECM and ECM-binding proteins (Fig. 3A). These included collagens and proteins associated with their synthesis (Col1a1/2, Col2a1, Col6a1/2, Col11a1/2, Col12a1, Pcolce), basal membrane-anchoring proteins (Prelp, Nid1), proteoglycans (Bgn, Fmod, Dcn, Vcan), glycoproteins (Sparc, Thbs1, Fn1, Mfge8), non-collagenous proteins (Comp), and chondrocyte proteins (Lect1, Clec3a), constituting a more fibrous tissue that is reflective of tensile strength, typical of the AF.
In addition to the 864 commonly expressed proteins, there were also proteins that were exclusively detected in either the NP or AF compartments that may provide additional insights (Fig. 3B; Table 1, Table 2). However, these were more variable between disc samples; thus, we imposed a filter to assess only proteins detected in over 50 % of the samples as the criterion for an exclusively expressed protein. Of the 1,617 proteins exclusively detected in the NP, 189 satisfied this criterion (Fig. 3B; Table 2). These proteins are represented by members of the solute carrier family (Slc2a1, Slc12a1, Slc44a2, Slc44a1), cytoskeleton (Dsp), gap junctions (Gja1), to lysosomal function (Atp1b3, Atp2b4, Atp6v1e1, Lamp1), that could be involved in protein clearance and coping with osmotic changes in the NP [35], [36], [37].
Table 1.
Partial list of proteins highly abundant or exclusively detected in AF categorized according to their biological functions annotated from Uniprot.
| Biological Function | Protein Name | Gene Symbol | Log2(Fold change) (AF/NP) | Exclusive in AF |
|---|---|---|---|---|
| ECM | ||||
| Collagens | ||||
| Collagen alpha-1(XII) chain | Col12a1 | 8.43 | ||
| Collagen alpha-2(I) chain | Col1a2 | 7.69 | ||
| Collagen alpha-1(I) chain | Col1a1 | 6.69 | ||
| Collagen alpha-1(XI) chain | Col11a1 | 4.51 | ||
| Collagen alpha-1(II) chain | Col2a1 | 4.62 | ||
| Collagen alpha-1(IX) chain | Col9a1 | 2.36 | ||
| Collagen alpha-1(VI) chain | Col6a1 | 3.94 | ||
| Collagen alpha-1(V) chain | Col5a1 | – | √ | |
| Collagen alpha-2(V) chain | Col5a2 | √ | ||
| Collagen alpha-1(XIV) chain | Col14a1 | – | √ | |
| Collagen alpha-2(IV) chain | Col4a2 | – | √ | |
| Proteoglycans | ||||
| Hyalectan | ||||
| Versican core protein | Vcan | 7.55 | ||
| SLRPs | ||||
| Biglycan | Bgn | 5.47 | ||
| Decorin | Dcn | 5.73 | ||
| Fibromodulin | Fmod | 8.11 | ||
| Lumican | Lum | 3.88 | ||
| Prolargin | Prelp | 4.95 | ||
| Mimecan | Ogn | – | √ | |
| Von Willebrand factor A family | ||||
| Matrilin-2 | Matn2 | – | √ | |
| Matrilin-3 | Matn3 | 4.83 | ||
| Matrilin-4 | Matn4 | – | √ | |
| Basement membrane | ||||
| Nidogen-1 | Nid1 | 2.21 | ||
| Nidogen-2 | Nid2 | 2.58* | ||
| Negative regulators of angiogenesis | ||||
| Thrombospondin-1 | Thbs1 | 4.21 | ||
| Thrombospondin-2 | Thbs2 | – | √ | |
| Leukocyte cell-derived chemotaxin 1 | Lect1 | 4.68 | ||
| Pigment epithelium-derived factor | Serpinf1 | 3.84 | ||
| Decorin | Dcn | 5.73 | ||
| Collagen alpha-2(IV) chain;Canstatin | Col4a2 | – | √ | |
| Involved in TGF-βsignaling | ||||
| Thrombospondin-1 | Thbs1 | 4.21 | ||
| Mimecan | Ogn | – | √ | |
| Protein-lysine 6-oxidase | Lox | – | √ | |
| Dermatopontin | Dpt | – | √ | |
| Fibromodulin | Fmod | 8.11 | ||
| Biglycan | Bgn | 5.47 | ||
| Decorin | Dcn | 5.73 | ||
| Mimecan | Ogn | – | √ | |
| Alpha-2-HS-glycoprotein | Ahsg | 3.42 | ||
| Tartrate-resistant acid phosphatase type 5 | Acp5 | 1.39 | ||
| Inorganic pyrophosphatase | Ppa1 | 0.63* | ||
| Osteopontin | Spp1 | 2.48* | ||
| Collagen alpha-1(IX) chain | Col9a1 | 2.37 | ||
| Leukocyte cell-derived chemotaxin 1 | Lect1 | 4.69 | ||
| Oxidative stress | ||||
| Extracellular superoxide dismutase [Cu-Zn] | Sod3 | 1.87* | ||
| Copper transport protein ATOX1 | Atox1 | 0.25* | ||
| Clusterin | Clu | 3.01 | ||
| AF markers | ||||
| Tenomodulin | Tnmd | – | √ | |
| Collagen alpha-1(V) chain | Col5a1 | – | √ | |
| Collagen alpha-2(V) chain | Col5a2 | – | √ | |
| Collagen alpha-1(XII) chain | Col12a1 | 8.43 | ||
| Leukocyte cell-derived chemotaxin 1 | Lect 1 | 4.69 | ||
| Proteins known to be associated with IVD disease | ||||
| Lysyl oxidase homolog 3 | Loxl3 | 1.17* | ||
| Cartilage intermediate layer protein 1 | Cilp | 2.70* | ||
| Apolipoprotein E | ApoE | 2.61 | ||
| Fibulin-5 | Fbln5 | – | √ | |
| Connective tissue growth factor | Ctgf | – | √ | |
| Cartilage oligomeric matrix protein | Comp | 2.69 | ||
*Not statistically significant after p-value adjustment for multiple testing.
Table 2.
Partial list of proteins highly abundant or exclusively detected in NP categorized according to their biological functions annotated from Uniprot.
| Biological Function | Protein Name | Gene Symbol | Log2(Fold change) (NP/AF) | Exclusive in NP |
|---|---|---|---|---|
| Cell-cell communication | ||||
| Cadherin-2 | Cdh2 | – | √ | |
| Gap junction alpha-1 protein | Gja1 | – | √ | |
| CD44 antigen | CD44 | – | √ | |
| Vinculin | Vcl | 1.82* | ||
| Desmoplakin | Dsp | – | √ | |
| Catenin alpha-1 | Ctnna1 | – | √ | |
| Alpha-actinin-4 | Actn4 | 2.60 | ||
| Transport | ||||
| Vesicle mediated transport | ||||
| T-complex protein 1 subunit theta | Cct8 | 2.62* | ||
| T-complex protein 1 subunit alpha | Tcp1 | 1.93 | ||
| Ras-related protein Rab-38 | Rab38 | – | √ | |
| Vesicle-associated membrane protein 3 | Vamp3 | – | √ | |
| Ras-related protein Rab-1B | Rab1b | – | √ | |
| Ras-related protein Rab-5C | Rab5c | – | √ | |
| Vesicle-trafficking protein SEC22b | Sec22b | – | √ | |
| Syntaxin-12 | Stx12 | – | √ | |
| Coatomer subunit beta | Copb2 | – | √ | |
| Coatomer subunit alpha;Xenin;Proxenin | Copa | – | √ | |
| Transmembrane emp24 domain-containing protein 9 | Tmed9 | – | √ | |
| Ras-related protein Rab-10 | Rab10 | – | √ | |
| Endocytosis | ||||
| Prolow-density lipoprotein receptor-related protein 1 | Lrp1 | 1.87* | ||
| Lysosomal-trafficking regulator | Lyst | – | √ | |
| Protein kinase C and casein kinase II substrate protein 3 | Pacsin3 | – | √ | |
| Clathrin heavy chain 1 | Cltc | 5.52 | ||
| EH domain-containing protein 4 | Ehd4 | – | √ | |
| AP-2 complex subunit beta | Ap2b1 | – | √ | |
| Protein kinase C and casein kinase substrate in neurons protein 2 | Pacsin2 | – | √ | |
| Lysosome related | ||||
| Lysosome-associated membrane glycoprotein 2 | Lamp2 | 1.88 | ||
| Lysosome-associated membrane glycoprotein 1 | Lamp1 | – | √ | |
| Ras-related protein Rab-5C | Rab5c | – | √ | |
| Ras-related protein Rab-7a | Rab7a | 2.61* | ||
| Vacuolar protein sorting-associated protein 35 | Vps35 | – | √ | |
| Choline transporter-like protein 2 | Slc44a2 | – | √ | |
| Cation-dependent mannose-6-phosphate receptor | M6pr | – | √ | |
| Glucose Transport | ||||
| Solute carrier family 2, facilitated glucose transporter member 1 | Slc2a1 | – | √ | |
| Protein kinase C and casein kinase II substrate protein 3 | Pacsin3 | – | √ | |
| Transporter | ||||
| Choline | ||||
| Choline transporter-like protein 2 | Slc44a2 | – | √ | |
| Fe2+/Fe3+ | ||||
| Melanotransferrin | Mfi2 | 1.90* | ||
| Sideroflexin-3 | Sfxn3 | – | √ | |
| Ceruloplasmin | Cp | 2.72* | ||
| Na+/Cl-/K+ | ||||
| Solute carrier family 12 member 2 | Slc12a2 | – | √ | |
| Ca2+ | ||||
| CSC1-like protein 1 | Tmem63a | – | √ | |
| Coupled to ATP activity | ||||
| Sarcoplasmic/endoplasmic reticulum calcium ATPase 1 | Atp2a1 | 0.43* | ||
| Calcium-transporting ATPase | Atp2b4 | – | √ | |
| V-type proton ATPase subunit G 1 | Atp6v1g1 | 1.14* | ||
| V-type proton ATPase subunit E 1 | Atp6v1e1 | – | √ | |
| V-type proton ATPase catalytic subunit A | Atp6v1a | 2.49* | ||
| Cell response to stress | ||||
| Heat shock protein HSP 90-beta | Hsp90ab1 | 4.62 | ||
| Heat shock protein HSP 90-alpha | Hsp90aa1 | – | √ | |
| Transitional endoplasmic reticulum ATPase | Vcp | 3.28 | ||
| Protein disulfide-isomerase A6 | Pdia6 | 1.24* | ||
| Heat shock 70 kDa protein 1A | Hspa1a | 2.03* | ||
| Calnexin | Canx | – | √ | |
| Protein disulfide-isomerase A4 | Pdia4 | – | √ | |
| Myoferlin | Myof | – | √ | |
| Serpin B6 | Serpinb6 | 2.47* | ||
| NP markers | ||||
| Keratin, type II cytoskeletal 8 | Krt8 | 6.63 | ||
| Keratin, type I cytoskeletal 19 | Krt19 | 5.51 | ||
| Annexin A3 | Anxa3 | 2.70 | ||
| Cadherin-2 | Cdh2 | – | √ | |
| CD44 antigen | CD44 | – | √ | |
| Carbonic anhydrase 3 | Ca3 | 4.09 | ||
| Solute carrier family 2, facilitated glucose transporter member 1 | Slc2a1 | – | √ | |
| CD109 antigen | CD109 | 4.14 | ||
| CD81 antigen | CD81 | – | √ | |
| Homeostasis maintenance | ||||
| pH regulation | ||||
| V-type proton ATPase subunit G 1 | Atp6v1g1 | 1.14* | ||
| V-type proton ATPase subunit E 1 | Atp6v1e1 | – | √ | |
| V-type proton ATPase catalytic subunit A | Atp6v1a | 2.49* | ||
| Carbonic anhydrase 2 | Ca2 | 3.04* | ||
| Carbonic anhydrase 9 | Ca9 | – | √ | |
| Osmosis and cell volume | ||||
| Heat shock 70 kDa protein 1A | Hspa1a | 2.03* | ||
| Serine/threonine-protein kinase WNK1 | Wnk1 | – | √ | |
| Solute carrier family 12 member 2 | Slc12a2 | – | √ | |
| Signaling pathways | ||||
| Wnt | ||||
| Caveolin-1 | Cav1 | – | √ | |
| Catenin beta-1 | Ctnnb1 | – | √ | |
| Catenin delta-1 | Ctnnd1 | – | √ | |
| High mobility group protein B2 | Hmgb2 | – | √ | |
| Protein G-Phospholipase | ||||
| Guanine nucleotide-binding protein G(i) subunit alpha-2 | Gnai2 | – | √ | |
| Guanine nucleotide-binding protein G(s) subunit alpha isoforms short | Gnas | – | √ | |
| Guanine nucleotide-binding protein G(i) subunit alpha-1 | Gnai1 | – | √ | |
| Inactive phospholipase C-like protein 1 | Plcl1 | – | √ | |
| Phosphatidylinositide phosphatase SAC1 | Sacm1l | – | √ | |
| Inositol 1,4,5-trisphosphate receptor type 3 | Itpr3 | – | √ | |
*Not statistically significant after p-value adjustment for multiple testing.
By comparison, only 28 proteins were exclusively detected in over 50 % of the AF samples (Fig. 3B; Table 1). These included ECM proteins (Col5a1/2, Postn and Lama2), SLRP Osteoglycin (Ogn), glycoprotein (Thbs2, Lox), tendon and ligament marker (Tnmd, and Mmp13). The dominance of the ECM in the AF is consistent with the important structural function of the AF, modulating mechanical properties in providing tensile strength in response to bending or torsion forces [5], [38], and the preservation of structural matrix integrity [39]. The role of collagen type XII (Col12a1) in the AF is not clear, but possibly related to structure [40], [41].
Enriched biological pathways confirm major structural roles for the AF
To gain further insights into biological processes in the NP and AF, a Gene Ontology (GO) term analysis was performed by combining the differentially expressed (Fig. 3A) and exclusively detected (Fig. 3C) proteins. Representative GO terms for this group of NP proteins include “cellular macromolecule localization” (FDR q = 3.23 × 10−40), “secretion” (FDR q = 1.68 × 10−37), “regulation of transport” (FDR q = 3.81 × 10−18), “cytoskeleton organization” (FDR q = 8.18 × 10−17) and “cell junction organization” (FDR q = 7.37 × 10−11) (Fig. 4B).
Fig. 4.
Mouse NP is enriched for lysosomal and transport proteins whilst AF is enriched for ECM. (A) Proteins with strong non-additivity (‘interactions’) between compartments and levels (red depicts lumbar, grey depicts tail). (B) GO term enrichment of DEPs higher in NP proteome, as compared with all AF. (C) GO term enrichment of DEPs higher in AF proteome, as compared with all NP. (D) Immunofluorescence for type XII collagen (Col12a1) in mouse discs at multiple developmental stages, showing higher expression at P10 in the inner AF, which shifts to higher expression in the outer AF with age. (E) Immunohistochemistry of type I collagen (Col1a1) in mouse disc at multiple developmental stages, showing higher expression in outer AF, particularly in younger IVD. (F) Immunohistochemistry of type II collagen (Col2a1) in mouse disc at multiple developmental stages, showing higher expression in the inner AF that is seen in P10 through to 8-week-old mice. (G) Immunohistochemistry of decorin (Dcn) in mouse disc at multiple developmental stages, showing inner AF has slightly higher expression at P10, but is highest in outer AF at 4 and 8 weeks of age. (For interpretation of the references to color in this figure legend, the reader is referred to the web version of this article.)
Consistent with the structural role for the AF, representative GO terms for this group of AF proteins were mostly of extracellular matrix (ECM) components and organization. These included the most prominent GO terms “extracellular structure organization” (FDR q = 2.23 × 10−48), collagen fibril organization (FDR q = 2.26 × 10−26) and skeletal development (FDR q = 1.28 × 10−18) (Fig. 4C), consistent with its structural function. Interestingly, “Circulatory system development” (FDR q = 2.39 × 10−14), “vasculature development” (FDR q = 8.31 × 10−14) and “blood vessel morphogenesis” (FDR q = 2.67 × 10−11) were also among the top GO terms, reflecting the blood supplies around the outer AF structure.
To validate the characteristic structural components in the AF, we assessed the expression of types I and II collagens, and decorin in early postnatal tail IVDs by immunohistochemistry (Fig. 4D). While expression of these ECM proteins has been reported [7]; here, we showed a dynamic change in postnatal growth. In the mouse, type II collagen was not detectable in the NP region but strongly localized to the cartilage endplate, the inner AF but not the outer at P10 (Fig. 4D). At 8 weeks, this pattern changed with postnatal growth with increased expression in the NP periphery, but decreased expression level in the inner AF and differences between inner and outer AF were less clear (Fig. 4D).
A similar trend was observed for type I collagen, which was highly expressed and localized to the outer AF at P10 but decreased with growth and lost demarcation between the outer and inner AF by 8 weeks (Fig. 4D). Whereas, for decorin, a known critical marker for tendon-like structures and their maturation [42], it was initially localized to the cartilage end plate, but significant levels were detected in the AF with growth, localized more to the outer AF (Fig. 4D). Type XII collagen was localized to the region of the OAF bordering the IAF of P10 mice, with increasing expression in both inner and outer AF with age (Fig. 4D). These dynamic expression and localization patterns of ECM components at different stages are consistent with the literature [43] and shows the maturation of the AF.
Vacuoles in NP cells are heterogeneous, some with potential lysosomal and lipid storage functions
The high cellularity of the mouse NP enabled in-depth assessments of the cells in vivo in a healthy young disc. As expected, cellular activities dominated the high ranking GO terms for enriched NP proteins; in particular, processes related to lysosomes, vacuoles and membrane transports (Fig. 4B). Large vacuolar structures are a characteristic of notochordal-like cells in the NP [44]. The function of these vacuoles is not clear, but a relationship with lysosome-related organelles was suggested [45].
The top two GO terms showed a range of proteins involved in the formation and trafficking of vesicles such as vesicle-associated membrane protein 3 (Vamp3), vesicle-trafficking protein Sec22b, coatomer subunit beta (Copb2), ras-related protein Rab-1B (Rab1b). Of interest are the lysosome-related proteins such as lysosome-associated membrane glycoprotein 1 and 2 (Lamp1 and Lamp2), regulator complex protein Lamtor1, and ras-related protein Rab-5C (Rab5c) (Fig. 4B). This prompted us to study the relationship of lysosomal activity with the existence of large vacuoles in the notochordal-like cells in the mouse NP.
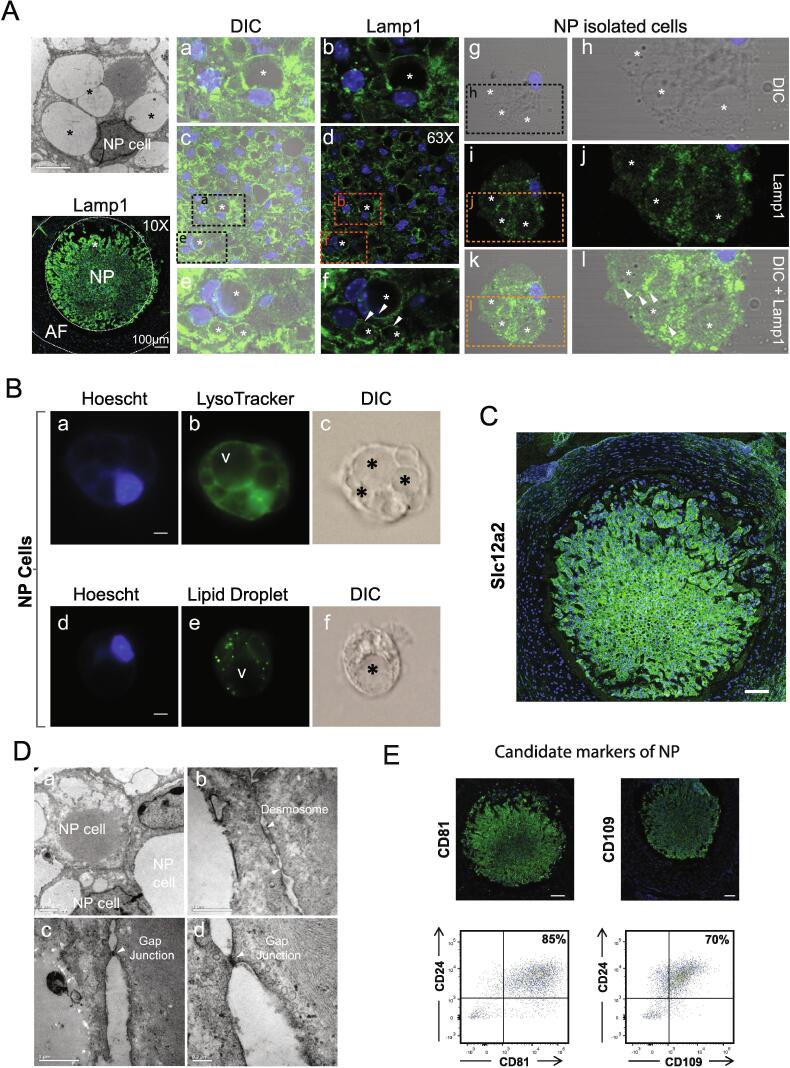
Ultrastructural analysis confirmed the presence of large vacuoles in most cells in the mouse NP (Fig. 5A). Lamp1 as a protein within the top two GO terms, specific to cells in the NP but not the AF (Fig. 5A). However, Lamp1 is localized to some small vacuoles/vesicles but not the large vacuoles within the cytoplasm of the NP cells, analyzed in frozen sections of the IVD, or primary culture of cells isolated from the NP (Fig. 5A). We estimated that 47.5 % (standard error 21.1 %) of all vacuoles/vesicles are Lamp1 positive (Supplementary Table S2). As the of vacuolar structure is preserved in isolated NP cells, this allowed the assessment of lysosomal organelles using the LysoTracker dye, showing distinct staining in the cell cytoplasm, but not within the large vacuoles (Fig. 5B). We further confirmed that there are some vacuoles containing lipid droplets in NP cells, but not localized large vacuoles (Fig. 5B). Thus, the characteristics of the vacuoles are heterogenous, with lysosomal or lipid storage functions, but the very large vacuoles are neither, with function yet to be determined.
Fig. 5.
Characterization of NP structures by transmission electron microscopy (TEM) and immunohistochemistry. (A) Transmission electron micrographs depict vacuole structures inside NP cells (labelled with white V) and immunofluorescence staining of NP cells show expression for lysosomal marker Lamp1. Lamp 1 is highly expressed in NP with presence of large vacuolar structures (IF and transmission electron microscopy) These structures resent with heterogenous staining for lysosomal marker Lamp 1 indicating potential existence of different types of vacuoles. (B) Microscopy images of live tail NP cells labelled with LysoTracker for 20 min (a) Hoescht, (b) LysoTracker and (c) differential interference contrast DIC and fixed tail NP cells labelled with LipidTOX green neutral lipid stain (d) Hoescht (e) LipidTOX and (f) DIC. (C) Immunofluorescence staining of IVD shows high expression of Slc12a2 in NP. (D) TEM of NP cells showing structures involved in cell–cell communication (a) shows presence of desmosome-like structures (b) as well as gap junctions (c & d, white arrows). (E) Immunofluorescence shows expression of Cd81 and Cd109 in NP. FACS analysis shows that Cd81 and Cd109 have high co-expression with NP marker Cd24. (For interpretation of the references to color in this figure legend, the reader is referred to the web version of this article.)
Cell-cell communication and role of ion channels in mouse NP cells
Enriched GO terms in the NP showed high levels of cell junction proteins, consistent with the high cellularity of the mouse NP and the presence of cell cluster contact (Fig. 4B). In parallel with a previous finding [46], adherens junction proteins such as N-cadherin (Cdh2) is an ‘exclusively detected” protein in the mouse NP (Fig. 3C; Fig. 4B). In addition, the enriched expression of anchoring junction proteins in the NP such as desmoplakin (Dsp), a desmosomal cadherin, and alpha-actinin-4 (Actn4), a component in the stability of F-Actin filaments support a high degree of cell–cell communication (Fig. 3A-C; Fig. 4B). Indeed, ultrastructural analysis of NP cells showed the presence of electron dense structures characteristic of anchoring junctions with a large adhesion plague from by aggregating desmosomes interacting with intermediate filaments of the cytoskeleton (Fig. 5D), as well as contacts consistent with adherens junctions formed via N-cadherins linking to F-actin filaments, and potential gap junctions via gap junction alpha-1 protein (Gja1) between the NP cells (Fig. 5D).
Another GO term-enriched in the NP is “regulators of transport” consisting of channel and transporter proteins across cell membranes such as choline transporter-like protein 1 and 2 (Slc44a1 and Slc44a2), iron transport sideroflexin-3 (Sfxn3), ceruloplasmin (Cp), ATPase subunit E1 (Atp6v1e1) and sodium potassium chloride transporters, solute carrier family 12 member 2 (Slc12a2/Nkcc1) (Fig. 3A-C; Fig. 4B). Slc12a2 is a member of the Na-K-Cl co-transporter proteins, and known to be involved in the response to hypertonic stress by regulating cell volume increase [47]. We showed that Slc12a2 is specifically localized to cells in the NP and not the AF (Fig. 5C). Further, a serine/threonine-protein kinase, Wnk1, regulates many ion co-transporters, and is a known upstream regulator of Slc12a2 [48], is also highly up-regulated in the NP (Fig. 3C; Fig. 4B). Wnk1 can be activated under hypertonic stress in a variety of cells. Our identification in NP cells would be consistent with the role of Wnk1 in regulating ion transporters such as Slc12a2 in NP cells in response to changes in hypertonicity and maintenance of cell volume [48].
Novel cell surface markers identified for NP cells
Specific cell surface markers are important for easy identification and purification of cells. Cd24 is a commonly used surface marker for NP cells [49], [50], however, it is not specific and expressed in multiple tissues [51]. Cell surface markers are best identified through a proteomic approach. With high cellularity of the mouse NP, we assessed for novel surface markers, and identified that Cd109 and Cd81 were highly or exclusively expressed in the mouse NP (Fig. 4B). Cd109 was noted as a proteomic marker for human NP cells, with antagonistic roles in the TGF-β pathway that promotes chondrogenic differentiation [7]. Immunostaining showed that both Cd109 and Cd81 were specifically expressed in the NP of the IVDs (Fig. 5E). FACs analysis of mouse NP cells for these markers that showed 85 % were double-positive for Cd24/Cd81 and 70 % for Cd24/Cd109 (Fig. 5E). Although Cd81 and Cd109 are expressed in other tissues, their relative specificity within the IVD context provided additional value for NP cell characterization and isolation.
Lumbar IVDs are enriched for mitochondrial proteins
Mouse IVD at the tail region is frequently used in studying disc biology and modelling degenerative states due to their ease of access, with an assumption that all IVD shared common biomechanical properties. However, this has not been thoroughly investigated. Interestingly, we identified significant differences in the protein profiles between tail and lumbar discs as whole IVD (NP + AF), as well as individual NP and AF compartments.
In the whole IVD assessment, there were 1,303 proteins commonly expressed between the lumbar and tail IVDs (Fig. 3D & E). Of these, 12 and 7 were statistically higher in lumbar and tail discs, respectively (Fig. 3D). There were also proteins that were exclusively detected in lumbar (3 8 0) or tail (1,031) discs; however, only 27 were detected in more than 50 % of the samples in lumbar discs and 1 in tail discs (Fig. 3E & F). Interestingly, the 27 lumbar-enriched proteins included an array of mitochondrial-related proteins (Acadl, Atp5j, Uqcrb, Uqcrfs1, Ndufs6, Gpd2, Cyc1, Ndufs3/4/9, Oxct1). With the exception of Oxct1, which is enriched in the lumbar AF, all are expressed higher in the lumbar NP (Fig. 3F). Proteins involved in neutrophil and eosinophil activities (Ltf, Mpo, Ccamp, Elane, and Prg2) were also enriched in the lumbar NP (Fig. 3F).
Next, to delineate lumbar to tail DEPs attributed to AF, NP, or as whole IVD (NP + AF), we separately ascertained lumbar to tail differences for proteins detected in the NP (Supplemental Fig. S5A) or AF (Supplemental Fig. S5B). The relationships are summarized in the Venn diagrams for proteins that are higher in the lumbar IVDs compared to tails (Fig. 3G), or higher in tail IVDs compared to lumbar (Fig. 3H) levels. There are a number of interesting observations. Of the 12 proteins that are detected higher in the lumbar as compared to AF (Fig. 3D, G), 6 were not attributed to the lumbar NP or AF when compared separately. That is, only the combined information between NP and AF provided a significant difference between lumbar and tail IVDs, but not individually; whereas the proteins higher in lumbar NP (4) and AF (15) (Supplemental Fig. S5A&B) are mostly distinct to the compartments, with only one protein shared between NP and AF (Fig. 3G). Consistently, these proteins are also enriched in mitochondria-related proteins; Sdha, Acaa2, Uqcrc1/2, Fh, Ndufs1, Aco2 and Etfb (Fig. 3G). Together with the lumbar-exclusive proteins set (Fig. 3F), the finding supports a higher mitochondrial activity in the lumbar IVDs, in both NP and AF compartments, but more so in the NP, suggesting a difference in the metabolic requirements.
There were 10 proteins “specifically” higher in the lumbar AF, including known AF markers Thbs1 and Cilp. On the other hand, proteins higher in the tail disc (Fig. 3H) were mostly ECM proteins (Collagens I/V/XV, Vcan and Fn1 for AF), and other proteins were related to skeletal cell differentiation, such as Serpine2 [52], Grem1 [53], S100a6 [54] and Rbm3 [55].
Proteins with spatial divergent expressing patterns between lumbar and tail IVDs
The distribution of proteins in the Venn diagram comparison of lumbar and tail IVD showed proteins that are specifically higher in either lumbar AF or NP, but not as a whole IVD (NP + AF) (Fig. 3G & H), raising the possibility of proteins with diverging or opposite trends between AF and NP compartments at the tail and lumbar regions, such that their combined (NP + AF) cancels out compartmental differences in the lumbar and tail level comparisons. To investigate this divergence, we fitted a two-way ANOVA model on IVD compartments and IVD levels with a statistical “interaction/non-additivity” behavior [56] for all the higher in lumbar (Fig. 3G) and tail (Fig. 3H) proteins, illustrated diagrammatically in Supplemental Fig. S6A & B.
We detected 22 proteins with moderate (P < 0.01 for the interaction term) non-additivity (Supplemental Fig. S6A). Six clearly (FDR < 0.05 for the interaction term) showed patterns of non-additivity (interactions) between compartments and levels (Fig. 4A), with 5 (Lamc1, Plec, Lamb2, Lyz2, and Capza2) displaying opposite expression trends between AF and NP at the lumbar and tail disc levels. The (4/5) majority of these have a high in the AF and low in the NP trend at the lumbar discs, but an opposite trend at the tail discs. Whereas expression of Lyz2 was low in lumbar AF and high on lumbar NP, with this trend reversed in tail discs. Thbs1, a known marker for AF and tendon-ligament, its expression is higher in the lumbar AF than in the NP; this trend remains but the difference is significantly reduced (Fig. 4A).
The known roles of these proteins ranged from the basal membrane (Lamc1, Lamb2), cytoskeletal protein (Plec), to monocyte/macrophage system (Lyz2). Of note, Lamc1 and Thbs1 are uniquely higher in lumbar AF but not in lumbar AF + NP (Fig. 3G). A full catalog of the trend lines for proteins uniquely higher in lumbar AF, lumbar NP and tail AF, tail NP can be found in Supplemental Fig. 6B, showing diverging relationships as well as parallel trends across the two IVD levels. The reason for these spatial divergent expression pattern for these set of proteins is not clear but may reflect the unique function of the mouse tail and/or its exposure to the less protected environment than the lumbar IVDs.
Non-varying proteins present an underlying common proteome of the IVD
In the DEP analyses, we noted a proportion of the detected proteins that do not vary with respect to compartment, level or gender. Through defined comparisons between datasets (Fig. 6A), we detected 181 such proteins, referred to as the “constant set”. GO-term analyses showed the majority of these proteins enriched for metabolic
Fig. 6.
The constant set of proteins represents the underlying scaffold of mouse IVD. (A) A set of proteins that did not change with respect to structure, level or gender was designated the ‘constant set’. Of the 277 that were detected, 181 of these were found in over 15 biological replicates, which consisted of matrisome and predominantly non-matrisome proteins. (B) Comparison of mouse and human ‘constant set’ showed 38 common proteins, which included proteoglycans Acan, Chad and Hapln1. (C) The expression levels (log2LFQ) of Acan are similar across all biological replicates. (D) Immunofluorescence of mouse IVD shows expression of Acan in both NP and AF.
processes, indicating basal cell function requirement that are important across all compartments of the IVD (Fig. 6B). Further, these cells, based on the GO terms are active in transcription and translation processes, as well as cellular metabolism needed for cell survival (Fig. 6B). A constant set was also identified in an atlas of the human IVD proteome, DIPPER [7], with 38 proteins that are common between human and mouse (Fig. 6C).
Of interest, there are 15 ECM/ECM-related proteins in the mouse constant set, none of which are collagens, suggesting different collagen types define IVD structural differences and function. Surprisingly, a number of proteoglycans were in this constant set, including aggrecan (Acan), a proteoglycan marker used to define the NP [57], that showed expression at comparable levels in the AF and NP (Fig. 6D, E), and detectable at similar levels by immunostaining in the mouse NP and AF tissues (Fig. 6F). As Acan was also detected as non-varying human dataset [7], we performed a detailed statistical analysis of the expression with Acan as a non-varying proteoglycan detectable across all IVD components and lumbar disc levels (Supplemental Fig. S7C). Thus, the proteomic findings in mouse and human seem to contradict established studies and a consensus that aggrecan expression is higher in the NP than AF [57].
To investigate this further, we examined Acan mRNA expression in recently published mouse (Supplemental Fig. S7A) and human transcriptomic data (Supplemental Fig. S7B) that further supported the lack of statistical differences in the Acan transcript between NP and AF in both the mouse and human IVDs.
Concordance between mouse transcriptome and proteome
The proteome reflects the accumulation of current and past transcriptional events. In expanding the information value of our proteomic data, we compared the protein expression to a recently reported mouse transcriptome dataset of the same stage (8 weeks) [18]. A direct sample-sample correlation heatmap showed a much higher correlation between the transcriptome and proteome in AF than in NP (Fig. 7A). A comparison of the difference between AF and NP in gene to protein expression showed a significant linear relationship (r = 0.41; p = 3.4 × 10−34) between the two data types (Fig. 7B).
Fig. 7.
Concordance between mouse transcriptome and mouse proteome, and comparison of mouse proteome with human disc proteome. (A) Correlation of mouse transcriptome and proteome show highest correlation for AF. (B) Differences between the mouse AF and NP transcriptome were correlated with the differences between AF and NP proteome and showed that NP had high expression of markers Krt8, Krt19, whilst AF was enriched for matrisome proteins including Col1a1/2, Col12a1, Vcan and Dcn. (C) Venn diagram showing the overlap between the total detected proteins in current mouse proteomic data and that in the young adult proteomic profiles in DIPPER (Tam, Chen et al. 2020). (D) Venn diagrams showing the overlaps among proteins detected in the AF and NP in current mouse proteomic data, and those in the AF and inner disc (NP and NP/IAF combined) compartments in DIPPER. A set of 1046 proteins was found to be detected only in mouse NP, whose GO enrichment was shown to the right. (E) Mouse and human proteome were compared and showed that human disc is most similar to mouse AF. (F) The differences between NP and AF in mouse and human were compared which showed high expression of KRT8 and KRT19 in NP, and enrichment of COL1A1 and COL12A1 in AF. (G) Venn diagram showing similarities and differences with mouse NP and human AF and NP. (H) Venn diagram showing similarities and differences with mouse AF and human AF and NP. (I) Venn diagram showing similarities and differences between the human ‘constant set’ with mouse NP and AF.
Of relevance are common gene/protein markers higher in the NP in both data types, including known NP markers (Krt8, Krt19), negative regulator of TGBβ (Cd109), cell cycle regulator Kctd12, transport proteins (Gstm2), cytoskeletal (Flnb, Tubb4b) and molecular chaperones (Hspb1). Common genes/proteins higher in the AF from both data types included ECM proteins such as Comp, Matn3, collagens (Col12a1, Col1a1/2, Col2a1, Col11a1, Col6a2), proteoglycans (Bgn, Dcn, Lum, Can, Fmod), glycoproteins (Sparc, Thbs1) and chondrocyte protein Lect1 (Cnmd) (Fig. 7B).
Molecular divergence and convergence between mouse and human IVDs
A clear understanding of the similarities and differences between mouse and human discs will inform on the relevance of the mouse disc biology to human. Using the DIPPER human disc proteomic dataset [7], we found 1166 proteins were detected in both human and mouse young adult IVD proteomes, representing 36.4 % of all proteins in both datasets (Fig. 7C). Further partitioning of these proteins into IVD compartments, we found 33 % of proteins (1046) exclusively detected in mouse NP (versus AF) (Fig. 7D). These proteins are mostly enriched in processes specific to mouse NP as compared with mouse AF (Fig. 7D; Fig. 4B), including “cellular macromolecular localization” and “secretion”, and “intracellular transport”, suggesting that the vacuole/lysosome-enriched notochordal-like cells may be more abundant in mouse than human NP, at approximately similar developmental stages (8wk vs 16yo).
We then performed sample-to-sample correlation analyses between proteomics profiles from murine and human discs. Murine AF proteome, regardless of tail or lumbar, is more closely related to the human AF than the NP proteome comparison (Fig. 7E & F). This is likely attributable to similar cell content and ECM structure of the AF. Whereas, the high cellularity and enrichment of notochordal-like cells in murine NP relative would expect to differ more to a human NP with cells sparsely embedded in ECM [19].
To better define the similarities and differences, we compared the DEPs in NP and AF of both species that revealed 4 common NP markers (KRT8, KRT19, CD109 and TNFRSF11B/OPG) (Fig. 7G), with Krt8/19/KRT8/19 as markers for the notochordal-like cells [15], [31], [32], [33], or cells of the chondro-osteogenic lineage (CD109) [34] and TNFRSF11B [58]). Comparison of the mouse and human AF and NP data revealed 19 and 7 common markers to AF and NP, respectively (Fig. 7H). The 19 proteins common to human/mouse AF are related to tendon/ligament or AF cell phenotype (LOX, TNMD, THBS2 and SLRPs). Next, we examined the expression patterns of DEPs or exclusively detected proteins (Fig. 3) higher in mouse NP or AF, in the human young adult proteomic data in DIPPER [7] (Supplemental Fig. 8). We found that some of the chondrocyte markers (COL2A1, MATN3/4, COL11A2, LECT2) that are enriched in the mouse AF are instead higher in human NP or inner disc (Supplemental Fig. 8A & C), suggesting that chondrocyte-like cells may exist in much lower quantities in the young adult NP of mouse (8wk) than human (16yo). Among the proteins enriched in mouse NP, only notochordal markers KRT8/19, CD109 and chondro-osteogenic marker TNFRSF11B remains higher in human NP (Supplemental Fig. 8B & D).
Lastly, we compared the mouse AF/NP DEPs with the human ‘constant set’ reported in DIPPER [7]. Here, we showed some proteins that are in the ‘constant’ in human, are DEPs in mouse. In fact, 6 % (13/216) and 14.3 % (29/203) of human constant proteins accounted for almost 20 % of the mouse AF DEPs (13/67) and 12.2 % of the NP DEPs (29/238), respectively (Fig. 7I). Together, while the signatures of AF in the two species remain largely conserved, clear molecular divergence can be observed between the mouse and human NP.
Discussion
The mouse as a model for IVD studies has supplemented our understanding of disc biology and related diseases [19]. However, relevance of mouse studies to human IVD and its degeneration is a subject of debate. Here, we provided a comprehensive proteomics dataset of “young and healthy” mouse IVDs covering major structures in the IVD (NP and AF), disc levels (lumbar and tail), and gender. We conducted extensive comparative assessments and validation, gaining further insights to mouse disc biology and its relevance to human IVD and disease states.
Cross-species variation is expected. Nonetheless, the AF bears the most structural similarity between mouse and human, with organized lamellar structures interspersed with cells. This is also reflected at the proteomic level between mouse and human, with tendon/ligament characteristics informing its function to resist tensile and torsional forces [7], [8]. We identified a characteristic matrisome for young and healthy mouse AF, which also appears to follow certain dynamic patterns with growth. Several markers display distinct demarcation between inner and outer AF, which diminishes with ageing, beginning as early as 4–8 weeks of age. For example, collagen XII was identified as a novel outer AF marker, with its expression localized to the inner AF at birth and then to the outer AF at 8 weeks of age. Although its function in the AF is not clear, previous studies suggest collagen XII has a regulatory role in the organization and mechanical properties of collagen fibril bundles in bone and connective tissues such as tendon, where their interactions with other ECM proteins result in the formation of ‘matrix bridges’, absorbing shear stresses from loading [59], and possibly facilitating cell–cell communication and cell organization [40]. Further, inactivation of Col12a1 in mice resulted in weakened muscle [60], disorganized collagen meshwork in bones [59], altered organization and impaired structure in tendon [40]. Together, this provides insights into matrix organization of the AF for its mechanical function, and orientation of cells within the AF lamellae.
Interestingly, proteins related to oxidative stress were elevated in the AF compared with the NP. This is consistent with an oxygen tension gradient radiating from the center of the disc, with the lowest levels of oxygen tension at the core [61]. The AF cells are likely to produce more reactive oxygen species (ROS) [62] as they may better utilize oxidative phosphorylation in a more oxygen-rich environment towards the outer AF. Clusterin (Clu) and Sod3 respond to extracellular ROS [63]. Oxidative stress can modify ECM by altering conformation of collagen molecules and thus increasing the loss of disc elasticity which is critical for structure maintenance [64]. It is possible that these stress proteins are in response to active turnover of ECM. For this, we can draw an analogy with the large and small diameter collagen fibers in tendons that are long-lasting or turnover rapidly, respectively [65]. In the context of the tendon, the small diameter collagen fibers serve to support the larger main fibers during loading, whereby damaged fibrils are broken down and replaced during rest, controlled by the circadian clock [65]. Whether such biological response occurs in the AF remains to be determined.
The mouse and human NPs, on the other hand, are less conserved. First, as compared with mouse AF, we detected higher cellularity in mouse NP, which is in contrast to the human NP with fewer cells and more ECM [7]. Mouse adolescent NP is dominated by vacuolated notochordal-like cells (NLC), whereas the human NP of the same stage is largely populated by chondrocyte-like cells. Nonetheless, there are studies indicating that mouse NPs do become less cellular with ageing, as demonstrated in inbred SM/J mice [17] and genetically modified strains such as premature ‘disc degeneration’ in mice with notochord-specific deletion of Ccn2, conditionally deleted Sox9 in Acan-expressing cells, or caveolin-1-null mice [66], [67], [68]. Further, in injury (IVD puncture) or mechanical-loading induced degeneration (tail looping) [67] mouse models for IDD, there is a loss of the vacuolated NLCs and a shift towards cells with chondrogenic and fibroblastic characteristics [26], which is also observed in human discs in ageing and degeneration [7], [8]. Thus, despite a comparable postnatal stage in terms of sexual maturity, the 8-week mouse and the 16yo human NPs are vastly different, and that the former may reflect a younger version of the latter.
In addition to the higher cellularity of NP in the mouse, the dominant expression of intracellular proteins in the NP could reflect specialized functions that differ from AF cells. Top cellular GO terms of DEPs in the NP implicate elevated activities for lysosomes, cell–cell communications, and gap-junctions. NLCs in the NP are known to contain large vacuoles [69]. Their composition and function, and questions as to whether they are vacuoles or vesicles, remain unresolved [70]. In zebrafish, vacuoles in notochord cell were shown to resemble lysosome organelles, expressing conventional lysosomal markers (Lamp1 and Lamp2), but the typical acidic environment was absent [45]. In young canine NP cells, these vacuole-like organelles were described as “membrane-limited structures derived from the plasma membrane which function in storage, transport or metabolism” that function as osmoregulatory elements and protect the cells from potential damaging swelling pressures [35]. Our study confirmed that Lamp1-expressing vacuoles are not acidic in healthy mouse NP cells. Whether these represent a novel class of lysosomes or they are lysosome precursors need further study. We show that the vacuoles in the mouse NP cells are heterogeneous, with some containing lipid droplets that could be related to energy storage, transport or metabolism [71]. There are also many larger vacuoles that are neither Lamp1 positive nor lipid droplet-containing. Whether these provide “hydration” or buffering function against compressive forces as some studies have suggested [35], [72], can be further investigated in the mouse.
The NP cells are continuously exposed to daily cycles of osmotic changes [73], [74], and can change ECM expression in cultured chondrocytes [36] and notochordal cells [37], with hyperosmotic conditions favoring a retention of notochordal cell phenotype [75]. The identification of Na-K-Cl transporters exclusively expressed in NP, including Slc12a1 and Slc12a2, and the kinase regulator Wnk1 at the molecular level reaffirms the need for NP cells to maintain ionic balance and cell volume in such an environment of dynamic osmolarity changes [47], [76]. Osmolarity stress responses involve early and late responses [77]. The early response corrects cell volume changes, and the late adaptive responses to hyperosmolarity were reported in chondrocytes [78] and endothelial cells [79]. Since cells in the IVD are non-excitable, the role of ion transporters/channels is not well investigated. Other studies have indicated a role in osmoregulation for acid-sensing ion channel 3 (Asic3) in rat NP cells [80], and aquaporins in human and canine NP [81]. Further, a comparative proteomic analysis and preliminary genetic assessment between the SM/J, a ‘poor healer’ strain with early-onset degeneration and LG/J, a ‘good healer’ strain protective for degeneration, showed differences in osmolarity regulating ion channels and osmotic stress response proteins [17], including a decrease in potassium channels (Slc12a2/4) in SM/J disc, known to regulate cell volume [47], [82].
While cells in the human NP are sparse, there are occurrences of cell clusters in healthy and degenerate discs. Their presence and role in human IVD biology is not well understood but can be modeled in the mouse NP. For example, consistent to previous studies, the proteome of mouse NP is enriched in Cdh2, shown to be critical in the maintenance of healthy NP cell phenotype and cell clusters [46]. In a study of human and porcine NP, CDH2 positive cells are needed to form cell clusters to maintain expression of NP-specific genes, linked to β-catenin-regulated signaling, and in NP cells isolated from degenerated discs to restore a healthy phenotype [46]. Further, involvement of anchoring junction proteins (Dsp, Actn4) identified in the current and other studies [69], highlight the importance of cell–cell communication in the formation and/or retention of cell clusters in NP homeostasis. Thus, a number of NP cell functions and characteristics can be modeled in the mouse for further investigation, including cell surface makers (Krt8, Krt19 and Cd24), and two novel cell surface markers (Cd109 and Cd81) that co-express with Cd24. Cd81 seems to be specific to the mouse NP cells, while Cd109 was reported in human NP cells [7].
Our current findings suggest while there are considerable similarities between tail and lumbar IVDs, distinct differences exist that need to be taken into consideration in study design and interpretation. In particular, cellular metabolism involving mitochondrial function, with significantly higher number of mitochondrial-related proteins were found in the lumbar IVD. It could be due to differences in oxygen tension and/or energy flux needed to control core temperature at the two levels of different environmental exposure. Tail IVDs, on the other hand, expressed higher levels of proteins (Grem1 and S100a6) associated with chondrogenic lineage [83]. A further puzzling finding is the presence of proteins that exhibit diverging or even opposing distribution trends from AF to NP, at the lumbar and tail IVD regions. This changing trend of disc compartment distribution between lumbar and tail discs is interesting. How this impacts biological function and relationship between mitochondrial and energy load in the mouse disc with ageing and degeneration warrants further investigation. Importantly, it uncovered potential differences, at least in the context of the mouse, that tail and lumbar IVDs are not necessarily equivalent. The relevance of these differences to disc biology, and whether they exist in the human setting, is yet to be determined, and investigators may need to consider this if caudal IVDs are suitable in their experimental design.
Our findings have several advantages over previous mouse IVD proteomics studies. Some studies have pooled the compartments [16] or used mice with a specific genetic alteration [17]. Here, we provided clear AF/NP compartmentalization from a normal mouse as a reference point. A limitation is that 8-week-old mice are not skeletally mature, an age where many mouse IVD studies are carried out. However, we elected to study the “adolescent” age of the mouse to reflect a healthy IVD, and for cross comparisons with an adolescent (16-year-old) human IVD in the DIPPER dataset [7].
In summary, this study provides a systematic and extensive proteomic landscape of the biology of a young and healthy mouse intervertebral disc, that helped identify molecular markers for the NP and AF structures, provided mechanistic insights to disc homeostasis and structural maintenance. Apart from confirming known structural roles of the AF, our findings revealed the involvement of lysosomes, osmoregulation, and cell–cell communication in the NP, critical for maintaining the hydrated status of the healthy discs. Further, we found differences between lumbar and tail IVDs in the mouse, and cross-species differences in the NP, as summarized in Fig. 8. It implies that the adolescent mouse NP may be a much younger version of the adolescent human NP, hence caution must be taken in the design and interpretation of data when using it as a model for human.
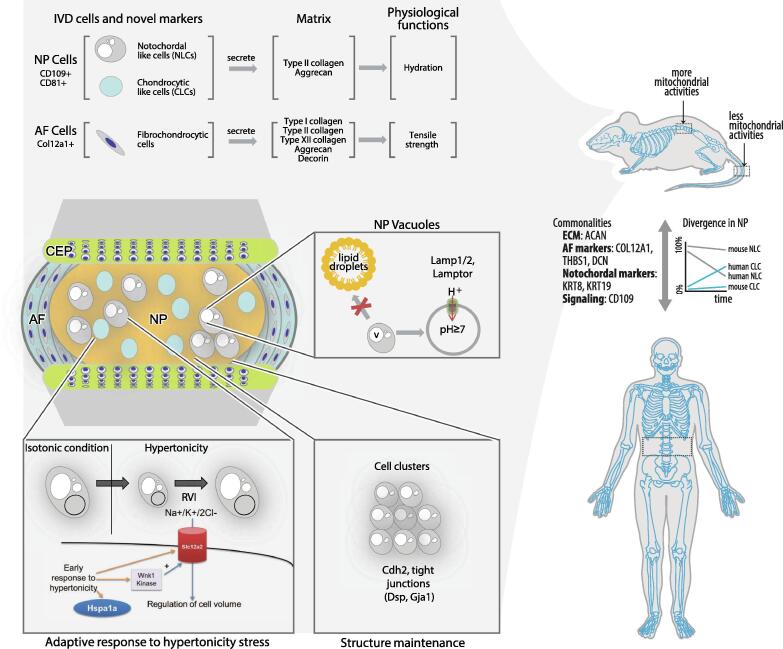
Fig. 8.
Schematic highlighting the key findings. Proteins associated with vacuoles and transport were identified, which may have implications in the regulation of hypertonicity stress in mouse NP, and a role for Cdh2 and tight junctions in cell clusters. Additionally, we identified additional potential markers Cd81 and Cd109. Comparison with human disc proteome revealed key commonalities and divergence which have implications in the use of mouse as animal models for studying the IVD.
Author contributions
MK designed experiments, performed sample preparation for mass spectrometry, immunohistochemistry, flow cytometry and TEM, data processing, analyses and interpretation, prepared figures and wrote the first draft of the mansucript. PKC performed bioinformatics analyses (ANOVA, PCA, SVM, DEPs, non-additivtiy, protein-RNA and human-mouse comparisons), data interpretation and prepared figures, developed the web interface, wrote and edited the manuscript. VT performed data analyses and interpretation, wrote and edited the manuscript. YZ performed sample preparation, immunohistochemistry, TEM and flow cytometry. OK performed sample preparation and immunohistochemistry. RS processed the raw data and was involved in data interpretation. TA provided the Foxa2mNE-Cre/ZEG mice for dissection and interpreted the data. KSEC, MKTT and WC provided critical input, interpreted results and were involved in writing. DC conceived ideas, supervised the project, interpreted data and results, and wrote the manuscript. DC and KSEC acquired funding to support this work. All authors contributed to editing and approved the manuscript.
Methods
IVD sample preparation, protein extraction
Animal licences were obtained from the Department of Health, Hong Kong, and animal ethics was approved by the Committee on the Use of Live Animals in Teaching and Research of The University of Hong Kong (CULATR 3956–16). Care of animals was in accordance with institution guidelines at the Centre for Comparative Medicine Research, which is an AAALAC International accredited service centre. To guide a visual and specific isolation of the NP and dissection of the AF, we used an in-house generated Foxa2mNE-Cre mouse model [26] crossed with the Z/EG reporter mouse to genetically tag notochord cells and descendants by the expression of the enhanced green fluorescent protein (eGFP). Eight-week-old Foxa2mNE-Cre/ZEG male and female mice were sacrificed and healthy tail and lumbar IVDs isolated and processed as independent samples. AF and NP were isolated from each IVD and proteins extracted using a chaotropic buffer (guanidine hydrochloride) as previously described [84] to enhance the solubilization of proteoglycans that are enriched in IVD tissues, maximizing total protein extraction.
Three biological replicates were harvested for each test condition (gender, levels and structures). In brief, samples were pulverized, sonicated on ice (5 s bursts with intensity at 40 % and 5 s breaks) before being resuspended into 15 volumes (v/w) of chaotropic buffer (pH 5.8) containing 50 mM NaAc, 4 M GdnHCl, 100 mM 6-aminocaproic acid, 5 mM benzamidine and 5 mM N-ethylmaleimide. Proteins were extracted at 4 °C for 48 h on an orbital shaker. Soluble proteins were collected after centrifugation at 13,200 g, 4 °C for 30 min, and pellets further extracted with the chaotropic buffer for another 24 h, prior to centrifugation. Soluble fractions were pooled, reduced with 4 mM DTT at 56 °C for 1 h, alkylated with 50 mM iodoacetamide at room temperature in the dark for 1 h, and reaction terminated by ethanol precipitation at 4 °C overnight. Proteins were recovered by centrifugation and pellets dried under vacuum and solubilized in 4 M urea, and protein content quantified using the BCA assay. 10 µg of proteins were subjected to sequential digestion using LysC (4 h) followed by trypsin (12 h) (Promega). Prior to LC-MS/MS analysis, peptides were dried under vacuum and desalted using C18 stage tips.
Mass spectrometry and data processing
Each sample was run in technical triplicates using an Orbitrap Fusion Lumos Tribrid Mass Spectrometer. Peptides were enriched on a 2 cm nano-trap column and further separated on a 50 cm long C18 column (ThermoFisher Scientific) at a flow rate of 300 nL/min delivered using Waters Acquity M−Class nano UPLC containing gradient from 100 % buffer A [5 % (v/v) acetonitrile in 0.2 % (v/v) formic acid] to 100 % buffer B [90 % (v/v) acetonitrile in 0.2 % (v/v) formic acid]. A total of 31 samples was collected (Supplemental Fig. S1A) and analyzed in triplicate, resulting in over 90 × 3 h LC-MS runs. The MS was controlled using Xcalibur 4.0 (ThermoFisher Scientific) and operated in a data-dependent acquisition mode. A full scan at 120,000 FWHM resolving power (at 200 m/z) was performed, followed by sequential HCD MS/MS scans at normalized collision energy of 31 and 30,000 FWHM resolution. The maximum injection times for both MS and MS/MS scans were 50 ms and 80 ms, respectively. Unassigned and single charged precursor ions were not selected for MS/MS with 45 s of dynamic exclusion for all fragmented peptides. The lock mass was enabled for accurate mass measurements using polydimethylcyclosiloxane (m/z, 445.12) ions.
Peptides were identified and quantified from raw mass spectrometric files using MaxQuant (v.1.5.3.30) [27]. Peak lists were generated for most intense MS peaks and extracted by MaxQuant. Database search was performed using the Andromeda search engine with the following parameters: against mus musculus Uniprot protein database (16,749 entries, May 2016) supplemented with sequences of contaminant proteins. Andromeda search parameters for protein identification were set to tolerance of 6 ppm for the parental peptide and 20 ppm for fragmentation spectra, and trypsin specificity allowing up to 2 mis-cleaved sites. Static modification of carboamidomethyl, the variable modifications of oxidation (M), and deamidation (NO) were specified. Minimal required peptide length was specified at 7 amino acids. Minimum of 2 unique peptides and proteins detected by at least 2 label-free quantitation (LFQ) ratio counts for protein identifications in one of the samples with a false discovery rate (FDR) of 1 % were considered as confident entries.
The ProteinGroups output files generated by MaxQuant analysis were processed as follow. The normalized label-free quantitation (LFQ) peptide intensity values, the numbers and percentage of sequence coverage of peptides (razor/unique) for each of the identified proteins, the MS/MS counts and intensities were imported into Perseus. For each of the analysis proteins identified only by site or by reverse database matching were excluded and remaining observed LFQ intensities were log2 transformed.
Functional bioinformatics analysis
The detected proteins were categorized into “matrisome” and “non-matrisome”, whereby “matrisome” were further divided into “core matrisome” and “non-core matrisome”, according to [28]. “Core matrisome” includes collagens, proteoglycans, and glycoproteins; whereas “non-core matrisome” includes “ECM affiliated”, “ECM regulators” and “secreted factors”. The mouse protein names were converted to human names using Ensembl’s BioMart (https://m.ensembl.org/info/data/biomart) before their corresponding gene families as categorized by HGNC (https://www.genenames.org/) were tabulated and ranked by descending order of frequencies.
To perform principal component analyses (PCA), we adopted an approached reported in DIPPER [7], by first selecting a smallest subset of proteins that maximally capture the detected values, before performing a missing value imputation by MICE [85]. Specifically, the number of missing values for protein were first ordered such that for all , where is the total number of detected proteins. A cutoff is chosen such , where F is a lagging factor (currently set to 200) designed to smoothen the slope estimations. PCA was performed using the singular value decomposition based R function prcomp() on the imputed data. SVM was trained and predicted suing the LIBSVM package [86].
To assess the relative importance of the three phenotypic factors on the proteome data, we performed linear regression for each of them on all or a subset of the detected proteins where is the log2(LFQ) expression for a protein and is one of those three phenotypic factors. The variance explained was estimated as where MSE stands for mean squared error and is the sample variance of protein .
To address: 1) cross-gender differences between male and female of the same IVD levels (lumbar or tail) and same IVD structures (NP or AF); 2) cross-level differences between lumbar to tail protein levels of the same IVD structure (AF or NP); and 3) cross-structure differences, we compared groups of protein profiles falling into such categories, by applying unpaired Student’s t tests to the transformed/normalized LFQ intensities for every protein. The individual p-values were converted to FDR (false discovery rate) q-values by the Holm method for more conservativeness. To control for false positives, we used an FDR method (FDR < 0.05). We ran 10,000 simulations of two samples for t-testing, one sample with 14 values from and one with 17 values from . By setting FDR < 0.05, we found 56% of the tests to be significant, i.e. achieving a power of 56%.
To assess the “non-additivity” between compartments and levels, we fitted a two-way ANOVA for each protein with an interaction term: , where is the log2(LFQ) of protein , , and . A protein was considered “moderately non-additive” if its two-way ANOVA FDR q-value is smaller than 0.05 and the p-value for interaction term is < 0.01. It is considered strictly non-additive if the FDR q-value of the interaction term is < 0.05.
Gene Ontology online tool, GOrilla [87] and GSEA (https://www.gsea-msigdb.org/) were used to perform a gene ontology (GO) for unique and differential proteins identified when comparing AF and NP. Only enriched GO terms with an FDR q-value < 10−2 were included for presentation. Human interpretation was involved in choosing representative GO terms for presentation.
Fluorescence immunohistochemistry of IVD tissues
8-week-old C57BL/6J (C57) mice were sacrificed and tail segments harvested, and immediately fixed with PBS containing 4 % (v/v) paraformaldehyde (Sigma, P-6148) for 16 h at 4 °C. Samples were then rinsed with PBS prior to removal of the ligaments. PBS containing 15 % (w/v) sucrose (Sigma, S8501) followed by 20 % (w/v) sucrose was used for cryo-preservation. Individual IVDs were then cut as one unit consists of the intervertebral disc and its two adjacent vertebral bodies and embedded in cryo-embedding media (OCT, 14020108926, Leica) orientated for sagittal or transverse-sectioning plane. Frozen blocks were sectioned at 8 μm, mounted on microscope slides (Art. J1800AMNZ, Menzel-Gläser, Superfrost® Plus, Thermo Scientific), and air-dried prior to staining.
For immunohistochemistry, tissues in each section was blocked at RT in PBS containing 3 % (w/v) bovine serum albumin (BSA, Sigma, A7906) followed by an overnight incubation at 4 °C with the respective primary antibodies (1:400 rabbit anti-Acan (Millipore, AB1031); 1:200 rabbit anti-Cd81 (ProSci, 5195); 1:200 rabbit anti-Cd109 (Abcam, ab203588); 1:800 rabbit anti-Col12a1 (Prof. Manuel Koch, University of Cologne, Germany); 1:200 rat anti-Lamp1 (Abcam, ab25245); 1:200 rabbit anti-Slc12a2 (Abcam, ab59791). Appropriate secondary antibodies conjugated with Alexa Fluor®488 (1:250) were used, and incubated for 20 min at room temperature. Finally, sections were washed, mounted (Vectorshield mounting medium for fluorescence, with DAPI, Vector, H1200), and imaged using an upright confocal microscope (Carl Zeiss LSM 710) connected to a digital microscope camera (AxioCam). Data were processed using ZEN v.5.5 SP1.
Electron microscopy
NP tissues were isolated, treated with 0.1 M cacodylate buffer (pH 7.2) containing 2.5 % (v/v) glutaraldehyde prior to fixation with 1 % (w/v) osmium tetroxide as previously described [88]. Cells were embedded in 2 % (w/v) agarose prior to treatment with uranyl acetate. Blocks were then dehydrated using graded series of ethanol and finally embedded in epoxy resin. Ultrathin sections were obtained using a Leica Ultramicrotome UCT (Leica, Microsystems Vienna, Austria) and collected on 400-mesh Formvar-coated copper grids. Sections were then stained for 45 min with aqueous uranyl acetate followed by lead citrate for 5 min.
Flow cytometry
Cells were obtained NP tissues from 8-week-old C57 mice digested for 1 h at 37 °C with 0.25 % (w/v) collagenase P (Roche) and 0.4 % (w/v) hyaluronidase (Merck Sigma) in HBSS. Isolated cells were fixed in 3.2 % (w/v) formaldehyde, blocked with 2% (w/v) BSA, and incubated with rabbit Cd109 (1:100) (Abcam, ab203588), rabbit Cd81 (1:50) (ProSci, 5195) or rat Cd24 (1:50) antibodies followed by incubation with appropriate secondary antibodies (1:200) (donkey anti-rabbit or donkey anti-rat) conjugated with Alexa 488 or Cyanine 3 (Cy3) respectively. Control unstained cells and single stained (Cd109 or Cd24) were used. NP cells were then analyzed by flow cytometry using a BD FACSCantoII Analyzer. Data were processed by FlowJo v10 (Tree Star, Ashland, OR).
Quantitative analysis of Lamp1 vacuoles
For the estimation of the percentage of Lamp1 positive and negative vacuoles, NP cells were isolated as described in previous section, fixed in 4 % (w/v) paraformaldehyde on a chamber slide followed by immunofluorescence staining of Lamp1 positive organelles by an overnight incubation using a Lamp1 antibody (1:200) (Abcam, ab25245). For quantitation, 5 regions from each well (n = 6) were imaged using confocal microscopy as previously described. 388 NP cells were analyzed and the total number of Lamp1 positive and negative vacuoles within the NP cells was determined.
Lysotracker and lipid droplet assays
Live tail NP cells were stained with 100 nM Lysotracker green DND-26 (Invitrogen, L7526) for 20 min according to the manufacturer’s instruction. For lipid stain, NP cells were first fixed in HBSS buffer containing 3.6 % (w/v) formaldehyde for 20 min, and then labelled using HCS LipidTOX™ green neutral lipid stain (1:1000) for 30 min (ThermoFisher Scientific, H34475) according to the manufacturer’s instruction. The fluorescent IHC images were captured with a fluorescent microscope (Olympus BX53) and processed (Olympus Cellsene Dimension 1.13).
Data and software availability
The RAW data has been deposited to the ProteomeXchange Consortium via the PRIDE [89] partner repository with the dataset identifier PXD006671. An interactive website hosting the data is available at: http://www.sbms.hku.hk/dclab/PRIMUS. The scripts for analyzing the data are available on https://github.com/hkudclab/mouse_ivd_proteomics.
Role of funding source
This work was in part supported by the Theme-based Research Scheme (TRS) (T12-708/12-N) of the Hong Kong Research Grants Council (RGC) awarded to KSEC (Project Coordinator and PI) and DC (co-PI), the RGC European Union - Hong Kong Research and Innovation Cooperation Co-funding Mechanism (E-HKU703/18) awarded to DC, and the RGC General Research Fund (GRF, 17126118) awarded to DC. We thank the Shenzhen “Key Medical Discipline Construction Fund” (No. SZXK077) for supporting PKC. All funding sources had no involvement in the study design, collection, analysis and interpretation of data, in the writing of the manuscript, nor in the decision to submit the manuscript for publication.
Declaration of Competing Interest
DC is serving on the editorial board of Matrix Biology. All other authors declare no competing interests.
Acknowledgments
We thank Dr. Rajkumar Ramalingam for performing the mass spectrometry and data processing.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.mbplus.2021.100082.
Appendix A. Supplementary data
The following are the Supplementary data to this article:
Supplementary figure 1.
Supplementary figure 2.
Supplementary figure 3.
Supplementary figure 4.

Supplementary figure 5.
Supplementary figure 6.
Supplementary figure 7.
Supplementary figure 8.
References
- 1.Adams M.A., Dolan P., McNally D.S. The internal mechanical functioning of intervertebral discs and articular cartilage, and its relevance to matrix biology. Matrix Biol. 2009;28(7):384–389. doi: 10.1016/j.matbio.2009.06.004. [DOI] [PubMed] [Google Scholar]
- 2.Vos, T., et al. (2012). Years lived with disability (YLDs) for 1160 sequelae of 289 diseases and injuries 1990–2010: a systematic analysis for the Global Burden of Disease Study 2010. 380(9859): p. 2163–2196. [DOI] [PMC free article] [PubMed]
- 3.Vo N.V. Molecular mechanisms of biological aging in intervertebral discs. J. Orthop. Res. 2016;34(8):1289–1306. doi: 10.1002/jor.23195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Chan W.C. Coming together is a beginning: the making of an intervertebral disc. Birth Defects Res. Part C: Embryo Today: Rev. 2014;102(1):83–100. doi: 10.1002/bdrc.21061. [DOI] [PubMed] [Google Scholar]
- 5.Heuer F., Schmidt H., Wilke H.-J. Stepwise reduction of functional spinal structures increase disc bulge and surface strains. J. Biomech. 2008;41(9):1953–1960. doi: 10.1016/j.jbiomech.2008.03.023. [DOI] [PubMed] [Google Scholar]
- 6.Cortes D.H., Elliott D.M. The Intervertebral Disc. Springer Vienna; Vienna: 2014. pp. 17–31. [Google Scholar]
- 7.Tam V. DIPPER, a spatiotemporal proteomics atlas of human intervertebral discs for exploring ageing and degeneration dynamics. Elife. 2020;9 doi: 10.7554/eLife.64940. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Yee A. Fibrotic-like changes in degenerate human intervertebral discs revealed by quantitative proteomic analysis. Osteoarthritis Cartilage. 2016;24(3):503–513. doi: 10.1016/j.joca.2015.09.020. [DOI] [PubMed] [Google Scholar]
- 9.Ranjani V. Proteomic analysis of degenerated intervertebral disc-identification of biomarkers of degenerative disc disease and development of proteome database. Glob. Spine J. 2016;6(S 01):p. WO025. [Google Scholar]
- 10.Maseda M. Proteomic analysis of human intervertebral disc degeneration. J. Nihon Univ. Med. Assoc.. 2016;75(1):16–21. [Google Scholar]
- 11.Rajasekaran S. Proteomic signatures of healthy intervertebral discs from organ donors: a comparison with previous studies on discs from scoliosis, animals, and trauma. Neurospine. 2020;17(2):426–442. doi: 10.14245/ns.2040056.028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Babu N.S. Quantitative proteomic analysis of normal and degenerated human intervertebral disc. Spine J. 2016;16(8):989–1000. doi: 10.1016/j.spinee.2016.03.051. [DOI] [PubMed] [Google Scholar]
- 13.Caldeira J. Matrisome profiling during intervertebral disc development and ageing. Sci. Rep. 2017;7(1):11629. doi: 10.1038/s41598-017-11960-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Cho H. Snapshot of degenerative aging of porcine intervertebral disc: a model to unravel the molecular mechanisms. Exp. Mol. Med. 2011;43(6):334–340. doi: 10.3858/emm.2011.43.6.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Sakai, D., et al. (2009). Differential phenotype of intervertebral disc cells: microarray and immunohistochemical analysis of canine nucleus pulposus and anulus fibrosus. Spine (Phila Pa 1976). 34(14): p. 1448-56. [DOI] [PubMed]
- 16.McCann M.R. Proteomic signature of the murine intervertebral disc. PLoS ONE. 2015;10(2) doi: 10.1371/journal.pone.0117807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Zhang Y. Early onset of disc degeneration in SM/J mice is associated with changes in ion transport systems and fibrotic events. Matrix Biol. 2018;70:123–139. doi: 10.1016/j.matbio.2018.03.024. [DOI] [PubMed] [Google Scholar]
- 18.Veras M.A. Transcriptional profiling of the murine intervertebral disc and age-associated changes in the nucleus pulposus. Connect Tissue Res. 2020;61(1):63–81. doi: 10.1080/03008207.2019.1665034. [DOI] [PubMed] [Google Scholar]
- 19.Alini M. Are animal models useful for studying human disc disorders/degeneration? Eur. Spine J. 2008;17(1):2–19. doi: 10.1007/s00586-007-0414-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Tam V. A comparison of intravenous and intradiscal delivery of multipotential stem cells on the healing of injured intervertebral disk. J. Orthop. Res. 2014;32(6):819–825. doi: 10.1002/jor.22605. [DOI] [PubMed] [Google Scholar]
- 21.McCann M.R. Tracing notochord-derived cells using a Noto-cre mouse: implications for intervertebral disc development. Dis. Model Mech. 2012;5(1):73–82. doi: 10.1242/dmm.008128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Siemionow, K., et al. (2011). The effects of age, sex, ethnicity, and spinal level on the rate of intervertebral disc degeneration: a review of 1712 intervertebral discs. Spine (Phila Pa 1976). 36(17): p. 1333-9. [DOI] [PMC free article] [PubMed]
- 23.Wang Y.X. Prevalence and sex difference of lumbar disc space narrowing in elderly chinese men and women: osteoporotic fractures in men (Hong Kong) and osteoporotic fractures in women (Hong Kong) studies. Arthritis Rheum. 2013;65(4):1004–1010. doi: 10.1002/art.37857. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.de Schepper, E.I., et al. (2010). The association between lumbar disc degeneration and low back pain: the influence of age, gender, and individual radiographic features. Spine (Phila Pa 1976). 35(5): p. 531–6. [DOI] [PubMed]
- 25.Teraguchi M. Prevalence and distribution of intervertebral disc degeneration over the entire spine in a population-based cohort: the wakayama spine study. Osteoarthritis Cartilage. 2014;22(1):104–110. doi: 10.1016/j.joca.2013.10.019. [DOI] [PubMed] [Google Scholar]
- 26.Au T.Y.K. Transformation of resident notochord-descendent nucleus pulposus cells in mouse injury-induced fibrotic intervertebral discs. Aging Cell. 2020;19(11) doi: 10.1111/acel.13254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Tyanova S., Temu T., Cox J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat. Protoc. 2016;11(12):2301–2319. doi: 10.1038/nprot.2016.136. [DOI] [PubMed] [Google Scholar]
- 28.Naba A. The extracellular matrix: Tools and insights for the “omics” era. Matrix Biol. 2016;49:10–24. doi: 10.1016/j.matbio.2015.06.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Wisniewski J.R. A “proteomic ruler” for protein copy number and concentration estimation without spike-in standards. Mol. Cell Proteomics. 2014;13(12):3497–3506. doi: 10.1074/mcp.M113.037309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Hunter C.J., Matyas J.R., Duncan N.A. Cytomorphology of notochordal and chondrocytic cells from the nucleus pulposus: a species comparison. J. Anat. 2004;205(5):357–362. doi: 10.1111/j.0021-8782.2004.00352.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Lee C.R. A phenotypic comparison of intervertebral disc and articular cartilage cells in the rat. Eur. Spine J. 2007;16(12):2174–2185. doi: 10.1007/s00586-007-0475-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Kerr, G.J., et al. Decoding the intervertebral disc: Unravelling the complexities of cell phenotypes and pathways associated with degeneration and mechanotransduction. in Seminars in cell & developmental biology. 2017. Elsevier. [DOI] [PubMed]
- 33.Minogue B.M. Transcriptional profiling of bovine intervertebral disc cells: implications for identification of normal and degenerate human intervertebral disc cell phenotypes. Arthritis Res. Ther. 2010;12(1):R22. doi: 10.1186/ar2929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Bizet A.A. The TGF-beta co-receptor, CD109, promotes internalization and degradation of TGF-beta receptors. Biochim. Biophys. Acta. 2011;1813(5):742–753. doi: 10.1016/j.bbamcr.2011.01.028. [DOI] [PubMed] [Google Scholar]
- 35.Hunter C.J. Osmoregulatory function of large vacuoles found in notochordal cells of the intervertebral disc. Mol. Cell. Biomech.: MCB. 2007;4(4):227. [PMC free article] [PubMed] [Google Scholar]
- 36.Wuertz K. Influence of extracellular osmolarity and mechanical stimulation on gene expression of intervertebral disc cells. J. Orthop. Res. 2007;25(11):1513–1522. doi: 10.1002/jor.20436. [DOI] [PubMed] [Google Scholar]
- 37.Chen J. Matrix protein gene expression in intervertebral disc cells subjected to altered osmolarity. Biochem. Biophys. Res Commun. 2002;293(3):932–938. doi: 10.1016/S0006-291X(02)00314-5. [DOI] [PubMed] [Google Scholar]
- 38.O'Connell G.D. Human internal disc strains in axial compression measured noninvasively using magnetic resonance imaging. Spine. 2007;32(25):2860–2868. doi: 10.1097/BRS.0b013e31815b75fb. [DOI] [PubMed] [Google Scholar]
- 39.McDONALD, J.A., D.G. Kelley, and T.J.J.T.J.o.c.b. Broekelmann (1982). Role of fibronectin in collagen deposition: Fab'to the gelatin-binding domain of fibronectin inhibits both fibronectin and collagen organization in fibroblast extracellular matrix. 92(2): p. 485-492. [DOI] [PMC free article] [PubMed]
- 40.Izu Y. Collagen XII mediated cellular and extracellular mechanisms regulate establishment of tendon structure and function. Matrix Biol. 2020 doi: 10.1016/j.matbio.2020.10.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Schonborn K. Role of collagen XII in skin homeostasis and repair. Matrix Biol. 2020;94:57–76. doi: 10.1016/j.matbio.2020.08.002. [DOI] [PubMed] [Google Scholar]
- 42.Robinson K.A. Decorin and biglycan are necessary for maintaining collagen fibril structure, fiber realignment, and mechanical properties of mature tendons. Matrix Biol. 2017;64:81–93. doi: 10.1016/j.matbio.2017.08.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Smith S.M. Topographical variation in the distributions of versican, aggrecan and perlecan in the foetal human spine reflects their diverse functional roles in spinal development. Histochem Cell Biol. 2009;132(5):491–503. doi: 10.1007/s00418-009-0623-z. [DOI] [PubMed] [Google Scholar]
- 44.Hunter, C.J., J.R. Matyas, N.A.J.J.o.a. Duncan (2003). The three‐dimensional architecture of the notochordal nucleus pulposus: novel observations on cell structures in the canine intervertebral disc. 202(3): p. 279–291. [DOI] [PMC free article] [PubMed]
- 45.Ellis K., Bagwell J., Bagnat M. Notochord vacuoles are lysosome-related organelles that function in axis and spine morphogenesis. J. Cell Biol. 2013;200(5):667–679. doi: 10.1083/jcb.201212095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Hwang P.Y., Jing L., Chen J., Lim F.-L., Tang R., Choi H., Cheung K.M., Risbud M.V., Gersbach C.A., Guilak F., Leung V.Y., Setton L.A. N-cadherin is key to expression of the nucleus pulposus cell phenotype under selective substrate culture conditions. Sci. Rep. 2016;6(1) doi: 10.1038/srep28038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Lang F. Functional significance of cell volume regulatory mechanisms. Physiol. Rev. 1998;78(1):247–306. doi: 10.1152/physrev.1998.78.1.247. [DOI] [PubMed] [Google Scholar]
- 48.Anselmo A.N. WNK1 and OSR1 regulate the Na+, K+, 2Cl− cotransporter in HeLa cells. Proc. Natl. Acad. Sci. 2006;103(29):10883–10888. doi: 10.1073/pnas.0604607103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Richardson S.M. Notochordal and nucleus pulposus marker expression is maintained by sub-populations of adult human nucleus pulposus cells through aging and degeneration. Sci. Rep. 2017;7(1):1501. doi: 10.1038/s41598-017-01567-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Fujita N. CD24 is expressed specifically in the nucleus pulposus of intervertebral discs. Biochem. Biophys. Res. Commun. 2005;338(4):1890–1896. doi: 10.1016/j.bbrc.2005.10.166. [DOI] [PubMed] [Google Scholar]
- 51.Fang X. CD24: from A to Z. Cell Mol. Immunol. 2010;7(2):100–103. doi: 10.1038/cmi.2009.119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Santoro A. SERPINE2 Inhibits IL-1alpha-Induced MMP-13 expression in human chondrocytes: involvement of ERK/NF-kappaB/AP-1 pathways. PLoS ONE. 2015;10(8) doi: 10.1371/journal.pone.0135979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Leijten J.C. GREM1, FRZB and DKK1 mRNA levels correlate with osteoarthritis and are regulated by osteoarthritis-associated factors. Arthritis Res. Ther. 2013;15(5):R126. doi: 10.1186/ar4306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Piltti J. Rho-kinase inhibitor Y-27632 and hypoxia synergistically enhance chondrocytic phenotype and modify S100 protein profiles in human chondrosarcoma cells. Sci. Rep. 2017;7(1):3708. doi: 10.1038/s41598-017-03958-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Jayasena C.S., Bronner M.E. Rbms3 functions in craniofacial development by posttranscriptionally modulating TGF-beta signaling. J. Cell Biol. 2012;199(3):453–466. doi: 10.1083/jcb.201204138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Alin A., Kurt S. Testing non-additivity (interaction) in two-way ANOVA tables with no replication. Stat Methods Med Res. 2006;15(1):63–85. doi: 10.1191/0962280206sm426oa. [DOI] [PubMed] [Google Scholar]
- 57.Sive J.I. Expression of chondrocyte markers by cells of normal and degenerate intervertebral discs. Mol. Pathol. 2002;55(2):91–97. doi: 10.1136/mp.55.2.91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Feng Z.Y. Osteoprotegerin promotes the proliferation of chondrocytes and affects the expression of ADAMTS-5 and TIMP-4 through MEK/ERK signaling. Mol Med Rep. 2013;8(6):1669–1679. doi: 10.3892/mmr.2013.1717. [DOI] [PubMed] [Google Scholar]
- 59.Chiquet M. Collagen XII: Protecting bone and muscle integrity by organizing collagen fibrils. Int J Biochem Cell Biol. 2014;53:51–54. doi: 10.1016/j.biocel.2014.04.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Zou Y. Recessive and dominant mutations in COL12A1 cause a novel EDS/myopathy overlap syndrome in humans and mice. Hum Mol Genet. 2014;23(9):2339–2352. doi: 10.1093/hmg/ddt627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Urban, J.P., S. Smith, and J.C. Fairbank (2004). Nutrition of the intervertebral disc. Spine (Phila Pa 1976). 29(23): p. 2700-9. [DOI] [PubMed]
- 62.Wunderli S.L. Tendon response to matrix unloading is determined by the patho-physiological niche. Matrix Biol. 2020;89:11–26. doi: 10.1016/j.matbio.2019.12.003. [DOI] [PubMed] [Google Scholar]
- 63.Kim J.H. Protective effect of clusterin from oxidative stress-induced apoptosis in human retinal pigment epithelial cells. Invest Ophthalmol Vis Sci. 2010;51(1):561–566. doi: 10.1167/iovs.09-3774. [DOI] [PubMed] [Google Scholar]
- 64.Scharf B. Age-related carbonylation of fibrocartilage structural proteins drives tissue degenerative modification. Chem. Biol. 2013;20(7):922–934. doi: 10.1016/j.chembiol.2013.06.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Chang J. Circadian control of the secretory pathway maintains collagen homeostasis. Nat Cell Biol. 2020;22(1):74–86. doi: 10.1038/s41556-019-0441-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Bedore J. Impaired intervertebral disc development and premature disc degeneration in mice with notochord-specific deletion of CCN2. Arthritis Rheum. 2013;65(10):2634–2644. doi: 10.1002/art.38075. [DOI] [PubMed] [Google Scholar]
- 67.Bach F.C. Increased caveolin-1 in intervertebral disc degeneration facilitates repair. Arthritis Res Ther. 2016;18:59. doi: 10.1186/s13075-016-0960-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Tsingas M. Sox9 deletion causes severe intervertebral disc degeneration characterized by apoptosis, matrix remodeling, and compartment-specific transcriptomic changes. Matrix Biol. 2020;94:110–133. doi: 10.1016/j.matbio.2020.09.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Hunter C.J., Matyas J.R., Duncan N.A. The functional significance of cell clusters in the notochordal nucleus pulposus: survival and signaling in the canine intervertebral disc. Spine. 2004;29(10):1099–1104. doi: 10.1097/00007632-200405150-00010. [DOI] [PubMed] [Google Scholar]
- 70.Wang F. Formation, function, and exhaustion of notochordal cytoplasmic vacuoles within intervertebral disc: current understanding and speculation. Oncotarget. 2017;8(34):57800. doi: 10.18632/oncotarget.18101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Thelen A.M., Zoncu R. Emerging roles for the lysosome in lipid metabolism. Trends Cell Biol. 2017;27(11):833–850. doi: 10.1016/j.tcb.2017.07.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Bagwell J. Notochord vacuoles absorb compressive bone growth during zebrafish spine formation. Elife. 2020;9 doi: 10.7554/eLife.51221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Nerlich A.G. Temporo-spatial distribution of blood vessels in human lumbar intervertebral discs. Eur. Spine J. 2007;16(4):547–555. doi: 10.1007/s00586-006-0213-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Johnson Z.I., Shapiro I.M., Risbud M.V. Extracellular osmolarity regulates matrix homeostasis in the intervertebral disc and articular cartilage: evolving role of TonEBP. Matrix Biol. 2014;40:10–16. doi: 10.1016/j.matbio.2014.08.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Spillekom S. Increased osmolarity and cell clustering preserve canine notochordal cell phenotype in culture. Tissue Eng. Part C Methods. 2014;20(8):652–662. doi: 10.1089/ten.TEC.2013.0479. [DOI] [PubMed] [Google Scholar]
- 76.McManus M.L., Churchwell K.B., Strange K. Regulation of cell volume in health and disease. N. Engl. J. Med. 1995;333(19):1260–1267. doi: 10.1056/NEJM199511093331906. [DOI] [PubMed] [Google Scholar]
- 77.Alfieri R.R., Petronini P.G. Hyperosmotic stress response: comparison with other cellular stresses. Pflügers Archiv-Eur. J. Physiol. 2007;454(2):173–185. doi: 10.1007/s00424-006-0195-x. [DOI] [PubMed] [Google Scholar]
- 78.Angelis E. Induction of betaine-γ-aminobutyric acid transport activity in porcine chondrocytes exposed to hypertonicity. J. Physiol. 1999;518(1):187–194. doi: 10.1111/j.1469-7793.1999.0187r.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Petronini P.-G. Induction of BGT-1 and amino acid system A transport activities in endothelial cells exposed to hyperosmolarity. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2000;279(5):R1580–R1589. doi: 10.1152/ajpregu.2000.279.5.R1580. [DOI] [PubMed] [Google Scholar]
- 80.Uchiyama Y. Expression of acid-sensing ion channel 3 (ASIC3) in nucleus pulposus cells of the intervertebral disc is regulated by p75NTR and ERK signaling. J Bone Miner Res. 2007;22(12):1996–2006. doi: 10.1359/jbmr.070805. [DOI] [PubMed] [Google Scholar]
- 81.Snuggs J.W. Aquaporin expression in the human and canine intervertebral disc during maturation and degeneration. JOR Spine. 2019;2(1) doi: 10.1002/jsp2.1049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Mobasheri A. Potassium channels in articular chondrocytes. Channels (Austin). 2012;6(6):416–425. doi: 10.4161/chan.22340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Worthley D.L. Gremlin 1 identifies a skeletal stem cell with bone, cartilage, and reticular stromal potential. Cell. 2015;160(1–2):269–284. doi: 10.1016/j.cell.2014.11.042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Önnerfjord P. Quantitative proteomic analysis of eight cartilaginous tissues reveals characteristic differences as well as similarities between subgroups. J. Biol. Chem. 2012;287(23):18913–18924. doi: 10.1074/jbc.M111.298968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.van Buuren S., Groothuis-Oudshoorn K. mice: Multivariate imputation by chained equations in R. J. Stat. Softw. 2011;45(3):1–67. [Google Scholar]
- 86.Chang C.-C., Lin C.-J. LIBSVM: a library for support vector machines. ACM Trans. Intell. Syst. Technol. 2011;2(3):1–27. [Google Scholar]
- 87.Eden E. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinf. 2009;10(1):48. doi: 10.1186/1471-2105-10-48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Kudelko M. Label-Free Quantitative Proteomics Reveals Survival Mechanisms Developed by Hypertrophic Chondrocytes under ER Stress. J Proteome Res. 2016;15(1):86–99. doi: 10.1021/acs.jproteome.5b00537. [DOI] [PubMed] [Google Scholar]
- 89.Vizcaíno J.A. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 2016;44(D1):D447–D456. doi: 10.1093/nar/gkv1145. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The RAW data has been deposited to the ProteomeXchange Consortium via the PRIDE [89] partner repository with the dataset identifier PXD006671. An interactive website hosting the data is available at: http://www.sbms.hku.hk/dclab/PRIMUS. The scripts for analyzing the data are available on https://github.com/hkudclab/mouse_ivd_proteomics.