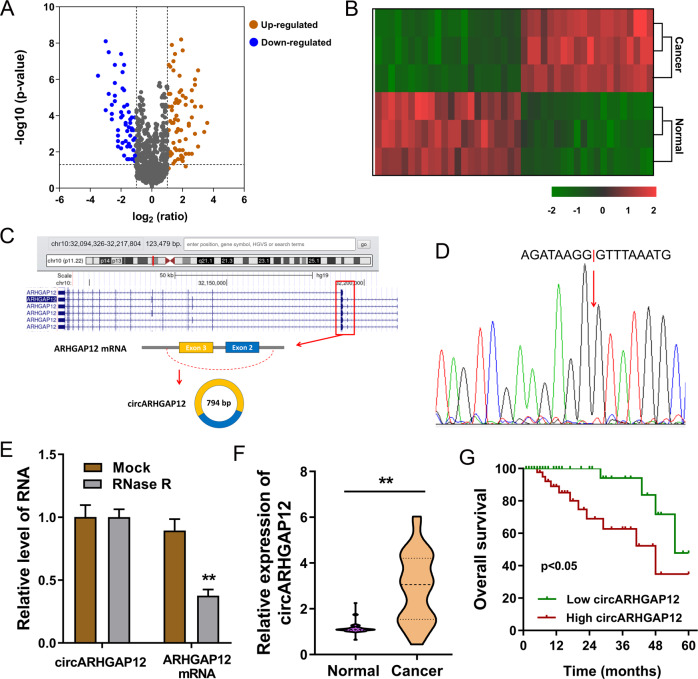
Fig. 1. circARHGAP12 was an unfavorable circRNA for cervical cancer.
A Volcano plot of circRNA microarray analysis revealed the des-regulated circRNAs within the cervical cancer tissue and adjacent normal tissue. B Heatmap of circRNA microarray analysis. C Schematic diagram demonstrated the generation of circARHGAP12 (hsa_circ_0000231, 796-bp) from exon-3 and exon-2 of ARHGAP12 gene through back splicing. D Sanger sequencing revealed the conjunction sites of circARHGAP12 back splicing. E RNA persistence testing using RNase R showed the expression of circular transcript (circARHGAP12) and linear mRNA (ARHGAP12 mRNA) detected using RT-PCR. F In the collected cervical cancer specimens’ group, circARHGAP12 expression was analyzed using RT-PCR. G Survival analysis using the Kaplan–Meier survival curve and the log-rank test demonstrated the survival rate of patients with higher or lower circARHGAP12 levels. P values were calculated by Student’s t-test. **p < 0.01.