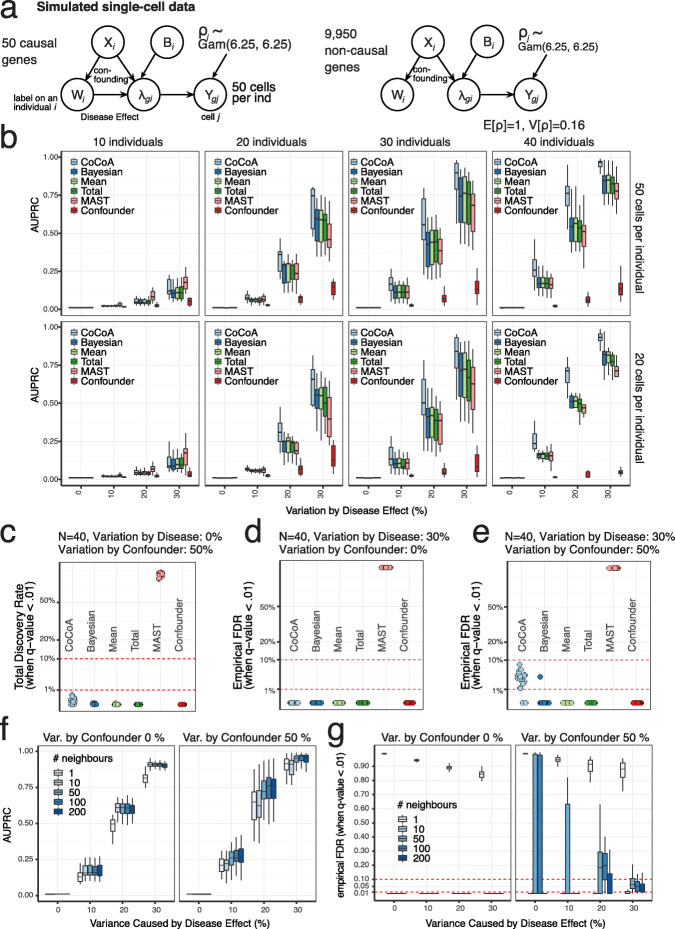
Fig. 2.
Simulation experiments. Extensive simulation experiments confirm that CoCoA-diff effectively adjusts existing confounding effects and improves statistical power of differential expression analysis. a Data generation scheme for simulation experiments. We simulate 50 causal and 9950 non-causal genes with or without disease-causing mechanisms (an edge between W and λ). Wi: disease label assignment for an individual i. Xi: confounding effects for an individual i. λgi: unobserved gene expression for a gene g of an individual i as a function of X and W. Ygj: realization of cell-level gene expression of a gene g with a cell j-specific sequencing depth ρj (stochastically sampled from Gamma distribution). Here, we simulated total five covariates consisting of confounding (X) and batch effect variables (B). b Simulation results when all the five covariates are confounding disease label assignment and gene expression values, accounting for 50% of mean expression variation (). Different subpanels correspond to different configurations of the number of individuals and cells per individual. Y-axis (AUPRC): area under precision recall curve (numerically integrated by DescTool [28] implemented in R); x-axis: the proportion of variation contributed by the disease label (). The following methods were considered: CoCoA: Wilcoxon’s ranksum test using individual-specific confounder-adjusted gene expression values δgi (the step 3 of Fig. 1c); Total: pseudo-bulk expression aggregated within each individual; Bayesian: Bayesian estimate of pseudo-bulk expression averaged over cells within each individual; Mean: pseudo-bulk expression averaged over cells within each individual; MAST: Model-based Analysis of Single-cell Transcriptomics [29] implemented in R (cell-level differential expression analysis); Confoudner: the estimated confounding effect μgi (the step 2 of Fig. 1c). c Total discovery rates of the differential expression methods when there were no disease effect. The fraction of positive discovery when multiple hypothesis-adjusted q-values were empirically calibrated by qvalue [30, 31] package controlled at 1% (y-axis). d Empirical false discovery rates of the differential expression methods when there were no confounding effect, but the 30% of individual-level expression variation is attributed to the disease effect (W → λ; ) on 50 causal genes. Y-axis: empirical false discovery rate, the frequency of the non-causal among genes with the estimated q-value below 0.01. e Empirical false discovery rates of the differential expression methods when there were substantial confounding effects on gene expressions () and the 30% of individual-level expression variation is attributed to the disease effect (W → λ; ) on 50 causal genes. Y-axis: empirical false discovery rate (the frequency of the non-causal among genes with the estimated q-value below 0.01); x-axis: different methods. f The performance of the CoCoA method with different settings of the k-NN parameters in the first matching step. Y-axis (AUPRC): area under precision recall curve (numerically integrated by DescTool [28] implemented in R); x-axis: the proportion of variation contributed by the disease label (). Variation by confounder: . g Empirical false discovery rates for the same experiments in f with different settings of the k-NN Parameter. Empirical false discovery rate: the frequency of the non-causal among genes with the estimated q value below 0.01