Fig. 2.
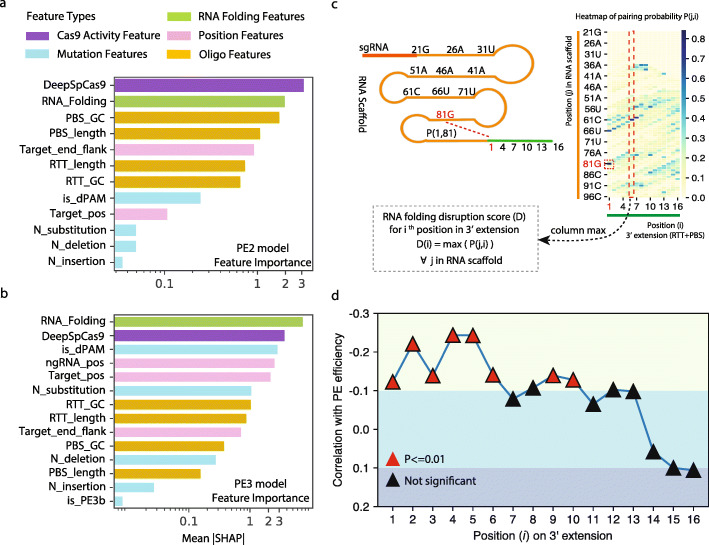
Features associated with PE efficiency. a, b Feature importance plot of the XGBoost regression model. Feature rankings are based on the mean absolute SHAP value for the PE2 and PE3 model. RNA folding features are combined for simplified visualization. Target_end_flank: number of nucleotides from target mutation to the end of RTT sequence. Target_pos: distance between target mutation and sgRNA nick site. ngRNA_pos: distance between ngRNA nick site and sgRNA nick site. c Schematic view of RNA-folding disruption score formulation. On the left, a pegRNA sequence consisting of an sgRNA (red), a scaffold sequence (orange), and an RTT sequence (green) is labeled with positions and nucleotides, such as 81G. The pairing probability between 81G and the first position in the RTT sequence is denoted as P(1,81). On the right is a heatmap of the pairing probability between each position in the scaffold and the 3′ extension sequence (i.e., RTT + PBS). P(1,81) is highlighted by a red dashed box. At bottom left, the formula to calculate D(i) is shown, where i represents the position in the 3′ extension. d Line plot showing the trend of correlations between the first 16 positions in the 3′ extension and the targeted editing frequency