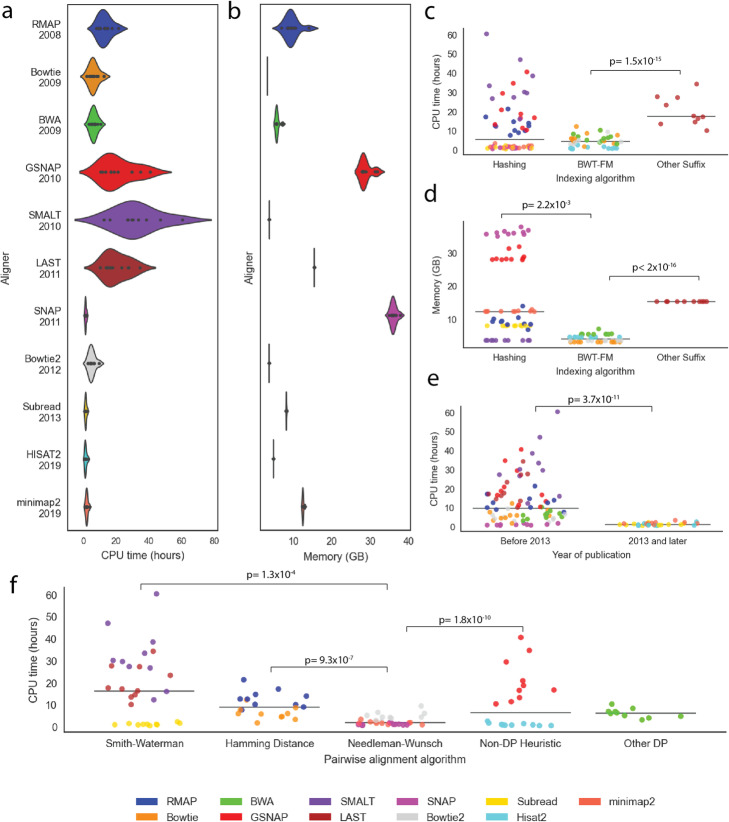
Fig. 4.
The effect of read alignment algorithms on the speed of alignment and computational resources. Results of the benchmarking performed on 11 surveyed DNA read alignment tools that can be installed through bioconda (RMAP, Bowtie, BWA, GSNAP, SMALT, LAST, SNAP, Bowtie2, Subread, HISAT2, and minimap2) additionally noted in Supplementary Table 2 and Supplementary Note 3. Each tool’s CPU time and RAM required were recorded for 10 different WGS samples from the 1000 Genomes Project. a, b Violin plots showing the relative performance (a CPU time and b RAM) of the benchmarked aligners. Aligners are ordered by year of release. c, d The relative performance (c CPU time and d RAM) of the benchmarked aligners grouped by the algorithm used for genome indexing and colored by individual aligners (BWT-FM CPU time vs. Suffix array CPU time: LRT, p value = 1.5 × 10−15, Hashing memory vs. BWT-FM memory: LRT, p value = 2.2 × 10−3, BWT-FM memory vs. Suffix Array memory: LRT, p value < 2 × 10−16). The legend of d is the same for c, e, and f. e The relative performance (CPU time) of the benchmarked aligners grouped by whether the tool was released before or after long-read technology was introduced (2013) and colored by individual aligners (LRT, p value = 3.7 × 10−11). f The relative performance (CPU time) of the benchmarked aligners grouped by the algorithm used for pairwise alignment and colored by individual aligners (Needleman-Wunsch CPU time vs. Smith-Waterman CPU time: Wald, p value = 1.3 × 10−4, Needleman-Wunsch CPU time vs. Hamming Distance CPU time: Wald, p value = 9.3 × 10−7, Needleman-Wunsch CPU time vs. Non-DP Heuristic CPU time: Wald, p value = 1.8 × 10−10)