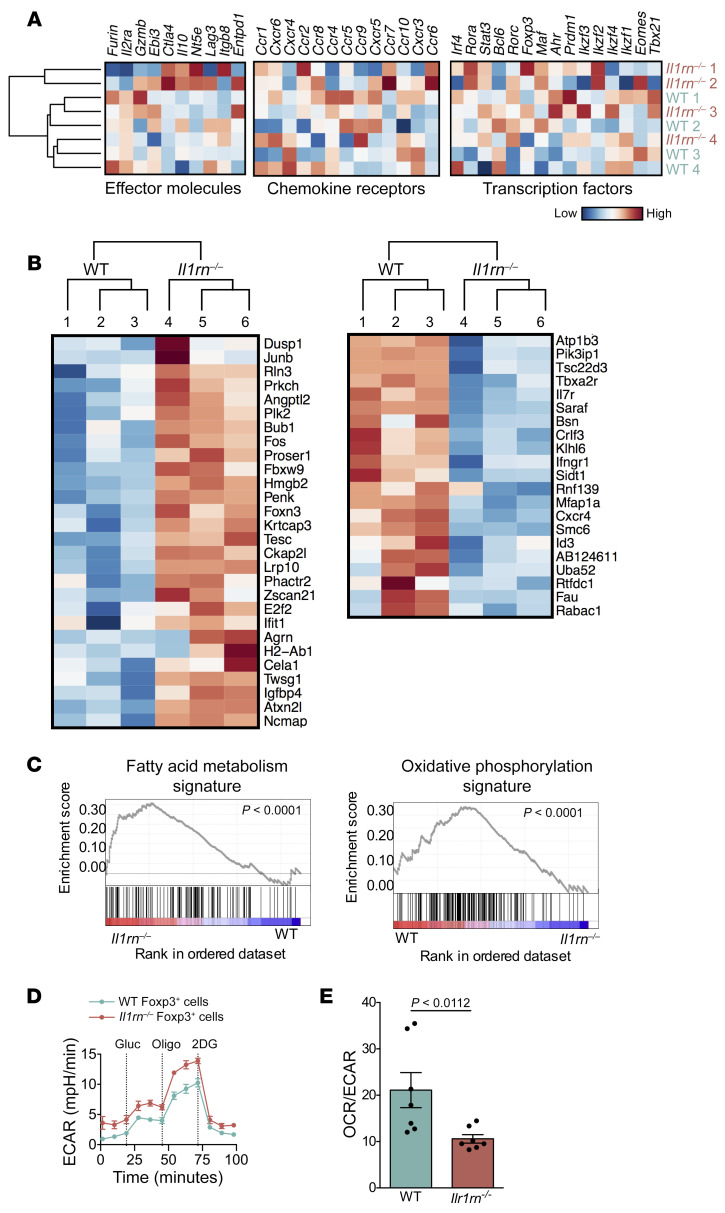
Figure 3. Il1rn–/– Tregs exhibit increased glycolysis.
(A and C) CD4+Foxp3eGFP+ Treg cells from WT and Il1rn–/– mice were sorted and RNA-seq transcriptomes were analyzed. (A) Systematic analysis of transcripts encoding the major functional molecules involved in Treg inhibitory function. Comparison is shown of normalized expression values of effector molecules, chemokine receptors, and transcription factors. (B) Heatmap of top 50 differentially expressed genes between CD4+Foxp3eGFP+ cells sorted from lymph nodes of 3 WT and 3 Il1rn–/– Foxp3eGFP animals. (C) Overexpression of fatty acid metabolism genes (left) and OxPhos genes (right) in Il1rn–/– vs. WT Tregs. (D and E) WT (1 × 105, n = 6) and Il1rn–/– (1 × 105, n = 6) lymph node CD4+Foxp3+ Tregs were equilibrated for 1 hour in unbuffered XF assay medium supplemented with 1 mM sodium pyruvate in an XF24 cell culture microplate. Extracellular acidification rate (ECAR) and oxygen consumption rate (OCR) were measured using an XF24 analyzer (Seahorse Bioscience). Compounds were injected during the assay at the following final concentrations in this glycolysis stress assay: 10 μM glucose, 1 μM oligomycin, and 50 μM 2-deoxyglucose (2DG). (D) ECAR (n = 12). (E) Basal OCR/ECAR ratio (n = 7). Data are expressed as mean ± SEM. Statistical significance was determined using 3-way ANOVA (D and E).